The general public can access information about accreditation and clinical sections.
Laboratory of Pathology
Laboratory of Pathology
The Laboratory of Pathology (LP) at the National Cancer Institute (NCI) is an integral component of the research and clinical community at the National Institutes of Health (NIH). Our goal is to be a globally recognized center of excellence in disease research, clinical diagnostics, and pathology education. The mission of the Laboratory of Pathology is to achieve the highest level of quality in research, diagnostics, and education.
Diagnostics
We take responsibility for Anatomic Pathology Services for the NIH as well as others who seek our expertise through consultation. Our goal is to provide the most accurate diagnostic interpretation in the most efficient manner, supported by a commitment to quality improvement.
Laboratory of Pathology Clinical Services
LP clinical and research staff can access information about LP's Clinical Services, using their NIH login credentials, via the internal portal here:
- Access LP Clinical Services (NIH login required)
This site includes information about: test menus and services provided; specimen collection and submitting guidelines; accreditation and regulatory compliance; quality management; safety and infection control; emergency management; and, technical and administrative policies and procedures.
Pathology Requisition Forms
PATHOLOGY OUTSIDE TISSUE EXAMINATION Form
NIH Methylation Profiling Consultation Requisition Form
NIH Methylation Profiling Consultation Requisition FormTissue Resource Committee (TRC) Requisition Form

Job Vacancies
We have no open positions in our group at this time, please check back later.
To see all available positions at CCR, take a look at our Careers page. You can also subscribe to receive CCR's latest job and training opportunities in your inbox.
News
Updates from the Laboratory of Pathology
Check back for occasional news from NCI's Laboratory of Pathology.
David Kleiner, MD, PhD receives the Hans Popper Hepatopathology Society Lifetime Achievement Award
Congratulations to Dr. David Kleiner for receiving the Hans Popper Hepatopathology Society’s Lifetime Achievement Award for outstanding contribution to Liver Pathology!
Congratulations Dr. Martha Quezado
Dr. Martha Quezado promoted to Co-Deputy Chief for the Laboratory of Pathology. Cited on LP's webpage...
Celebrating CCR Careers: David D. Roberts, Ph.D.
For more than forty years at NIH, David D. Roberts, Ph.D., has explored cellular processes regulated by the protein thrombospondin-1 in vascular and immune cells and developed new therapeutic approaches to treat cancers by shutting off the function of one of its receptors. (Read more)...
NCI Laboratory of Pathology Launches Bethesda Classifier Website - Methylscape
The Clinical Methylation Unit of the Laboratory of Pathology's NCI COMPASS program is excited to announce the release of our new website dedicated to the classification of central nervous system (CNS) tumors using DNA methylation data from Illumina methylationEPIC microarrays.
Developed by the neuropathology group of the Laboratory of Pathology at the National Cancer Institute (NCI), the Bethesda classifier is an advanced algorithm designed to enhance and expand upon the existing capabilities of the Heidelberg classifier. Our classifier includes the latest tumor entities, offering a more comprehensive tool for CNS tumor classification.
Key Features:
• Cutting-Edge Algorithm: The Bethesda classifier utilizes state-of-the-art techniques to provide precise and reliable CNS tumor classifications.
• Comprehensive Coverage: Includes newer tumor entities additional to the by the Heidelberg classifier, ensuring up-to-date diagnostic support.
• User-Friendly Interface: Our website offers an intuitive interface for researchers and clinicians to easily upload and analyze methylation data. Users are provided with the classification and Umap embedding.
We invite researchers, clinicians, and all interested parties to explore our new website and take advantage of the Bethesda classifier for their tumor classification needs.
Visit our website today to learn more and get started: https://methylscape.ccr.cancer.gov/
CC Grand Rounds John Laws Decker Lecture -
David E. Kleiner, MD, PhD - Distinguished Clinical Teacher Awardee 2024 Award
You are cordially invited to this reception, to celebrate Dr. David E. Kleiner as the 2024 Distinguished Clinical Teacher Awardee
When: Wednesday June 12th at 1-2pm
Where: Medical Board Room
To RSVP please sign up here by Friday May 31
The reception is after his 12pm CC Grand Rounds Decker – DCTA Lecture in Lipsett
Steatotic Liver Disease: From First Description to First Drug Approval Through The Eyes Of The Pathologist
If you have questions, contact Rita Stevens at rita.stevens@nih.gov
New AI tool classifies brain tumors using images of tumor slides
A new artificial intelligence (AI) tool called DEPLOY can analyze images of brain tumors to predict their methylation state — essentially which genes are turned “on” or “off” — and infer the tumor subtype with 95% accuracy. The tool, which could help scientists more easily and accurately identify brain tumor subtypes, appeared May 17, 2024, in Nature Medicine.
Dr. Jaffe elected to the Association of American Physicians
Elaine S. Jaffe, M.D., NIH Distinguished Investigator in the Laboratory of Pathology, was elected as a 2024 member of the Association of American Physicians.
Clinical Residency 1st Quarter 2024 News
Dr. Dilara Akbulut receives ISUP Award March 24, 2024
Congratulations Dr. Dilara Akbulut for receiving an International Society of Urological Pathology (ISUP) Stipend Award in recognition of excellent uropathologic research.
Celebrating CCR Careers: William G. Stetler-Stevenson, M.D., Ph.D.
For over three decades, William G. Stetler-Stevenson, M.D., Ph.D., has been a leader in the field of tissue inhibitors of metalloproteinases (TIMP) biology and extracellular matrix regulation of cancer. His team discovered that TIMPs have metalloprotease-independent activities and can affect cellular proliferation directly. (Read more)...
CCR researchers receive an array of HHS awards
In September 2023, three CCR researchers were honored with Departmental Awards from the Department of Health and Human Services (HHS). These awards celebrate scientists and administrators for their exceptional contributions to the CCR and HHS mission.
Elaine S. Jaffe, M.D., NIH Distinguished Investigator in the Laboratory of Pathology, received the Secretary’s Award for Meritorious Service. This award recognizes HHS employees for their achievements and for inspiring others in the field to improve their own performance. Jaffe is being honored for her pioneering advances in the diagnosis and treatment of patients with hematological neoplasms. She revolutionized the integration of traditional pathological methods with immunologic and genomic approaches, improved diagnosis methods for lymphoma and leukemia, and described multiple new disease entities. Her work has advanced clinical practice and management for numerous patients with blood cancers. (Read more)...
Lifetime Achievement Award: Elaine S. Jaffe, MD
The FASEB Excellence in Science Lifetime Achievement Award recognizes Elaine Jaffe’s pioneering work in the fields of hematology and hematopathology. Her work over the past 50 years has changed the way in which the diagnosis of lymphoma is made worldwide. As Distinguished Investigator at the National Cancer Institute (NCI), she is the final arbiter for challenging diagnostic problems submitted to her from around the world, personally reviewing more than 2,000 cases annually.
The award also recognizes Jaffe as a devoted educator and mentor. She has been recognized numerous times for her excellent teaching in the clinical setting and the laboratory. She has spoken at almost every major meeting in pathology, hematology, and oncology. She consistently publishes with trainees as the first author of her manuscripts. Jaffe provides guidance to these trainees on how to perform clinical research and how high a bar is required to publish meaningful results. In 2001, she won NCI’s Outstanding Mentor Award and in 2008 she received the Chugai Award for Excellence in Mentoring and Scholarship from the American Society for Investigative Pathology, a FASEB member society of which she is a member.
“Dr. Jaffe has had an illustrious career in which she has made multiple contributions to our understanding of lymphomas; redefined their classification in a way that can provide a useful guide for further research and for therapy; and has mentored and promoted the careers of numerous young hematopathologists,” says one of her nominators for the award, Robert Yarchoan, MD, at the National Institutes of Health.
2023 NCI Staff Scientists and Staff Clinicians Outstanding Mentor Award: David Peeney, Ph.D.
David Peeney, Ph.D. Staff Scientist in Bill Stetler-Stevenson’s group is the recipient of the 2023 NCI Staff Scientists and Staff Clinicians Outstanding Mentor Award. This is truly noteworthy as a contribution to training our next generation of scientists. He will be recognized at the 19th Annual Staff Scientist and Staff Clinician Retreat on Monday, April 24, 2023.
Laboratory of Pathology staff co-authors of December 14, 2022, Nature article “SARS-CoV-2 infection and persistence in the human body and brain at autopsy”. Dr. David Kleiner, Dr. Stephen Hewitt, Dr. Stefania Pittaluga, Dr. Martha Quezado and Mr. Kris Ylaya.
Coronavirus disease 2019 (COVID-19) is known to cause multi-organ dysfunction1,2,3 during acute infection with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), with some patients experiencing prolonged symptoms, termed post-acute sequelae of SARS-CoV-2 (refs. 4,5). However, the burden of infection outside the respiratory tract and time to viral clearance are not well characterized, particularly in the brain. Here we carried out complete autopsies on 44 patients who died with COVID-19, with extensive sampling of the central nervous system in 11 of these patients, to map and quantify the distribution, replication and cell-type specificity of SARS-CoV-2 across the human body, including the brain, from acute infection to more than seven months following symptom onset. We show that SARS-CoV-2 is widely distributed, predominantly among patients who died with severe COVID-19, and that virus replication is present in multiple respiratory and non-respiratory tissues, including the brain, early in infection. Further, we detected persistent SARS-CoV-2 RNA in multiple anatomic sites, including throughout the brain, as late as 230 days following symptom onset in one case. Despite extensive distribution of SARS-CoV-2 RNA throughout the body, we observed little evidence of inflammation or direct viral cytopathology outside the respiratory tract. Our data indicate that in some patients SARS-CoV-2 can cause systemic infection and persist in the body for months. Read more...
Dr. Zied Abdullaev participates in NCI-Connections Blog, “Diagnosing CNS Tumors More Precisely with Methylation Marks”.
An accurate diagnosis is the first critical step to ensure the best treatment for rare brain and spinal cord tumors. It enables treatment that is based on the unique biology of each cancer. Pathologists typically classify cancers by examining tumor tissue under a microscope. Doctors also rely on molecular marker tests. These can show changes in specific proteins or genes that are known to drive tumor growth and help guide treatment decisions. A novel way to classify central nervous system (CNS) tumors more precisely is by looking at patterns of chemical methyl marks on the DNA of cancer cells. This is known as DNA methylation. The presence or absence of methyl marks across the genome, which includes all your DNA, affects how active the underlying genes will be. Read more...
Dr. Elaine S. Jaffe receives the HHS Career Achievement Award
Elaine S. Jaffe, M.D., NIH Distinguished Investigator in the Laboratory of Pathology, was awarded a 2022 Career Achievement Award by the Department of Health and Human Services. These awards celebrate scientists and administrators for their exceptional contributions to the CCR mission.
Dr. Jaffe is recognized for her groundbreaking advances in the diagnosis and treatment of patients with blood cancers. With over 50 years of service to the NCI, she has worked to improve the outcome for patients with lymphoma and related diseases both at the NCI and internationally.
Dr. Jaffe has changed the way the diagnosis of lymphoma is made on a worldwide basis, 1) revolutionizing the integration of traditional pathological methods with immunologic and genomic approaches; 2) harmonizing for the first time the diagnosis of lymphoma and leukemia internationally; and 3) describing multiple new disease entities that have resulted in changes to clinical practice and management for numerous patients.
Whole Exome RNA Sequencing Now Available in the NCI-COMPASS, Laboratory of Pathology
November 30, 2021
The Comprehensive Oncologic Molecular Pathology and Sequencing Service (COMPASS), housed in CCR’s Laboratory of Pathology, is pleased to announce the implementation of CLIA whole-exome (all protein-coding genes) RNA-sequencing for CCR patients.
The sequencing pipeline will interrogate RNAseq of ~19k protein-coding genes and is primarily designed for the detection of established cancer-associated gene fusions (see fusion gene list at https://ccrod.cancer.gov/confluence/display/CCRLP/Molecular+Pathology+Test+Directory) . The pipeline will also represent a resource for research investigations to identify novel/rare gene fusions, provide gene expression data, as well as other applications. Detailed information about tumor/normal whole exome sequencing consenting, ordering and obtaining results may be found here https://ccrod.cancer.gov/confluence/pages/viewpage.action?pageId=73203825#CCRPolicies/StandardOperatingProcedures(SOPs)-ADCR. This test will be routinely run as part of the CRIS TSO-500 order of solid tumors. Going forward, a CRIS TSO-500 order for solid tumor testing will deliver a report describing key fusions identified from whole-exome RNA-sequencing, in addition to the previously available DNA sequencing assay encompassing >500 cancer genes. Please contact Dr. Liqiang Xi for questions.
As a reminder, all CCR clinical research operations SOPs may be found on the Office of the Clinical Director’s clinical standard operating procedures page and can also be accessed through the CCR Central Conducting Clinical Trials section.
As part of our efforts to enable you to apply and develop new precision medicine approaches, CCR established COMPASS to provide state of the art CLIA-certified clinical sequencing services.
The availability of whole exome RNA-sequencing adds yet another tool in the armamentarium of clinical genomic testing for our cancer patients and is an important step in CCR’s precision medicine activities.
Dr. David Kleiner's "At NCI, on the trail of drug-induced liver injury" featured in CAP Today
February 28, 2020
When looking at a liver biopsy, always suspect there might be drug-induced liver injury. “But then try hard to prove there isn’t. It’s always a diagnosis of exclusion and pattern evaluation,” says David E. Kleiner, MD, PhD, reference pathologist for the Drug-Induced Liver Injury Network. In that role, he pieces together clues that point to a drug having injured a liver—or not.
Drug-induced liver injury is rare, and few pathologists see it. “The incidence is between one in 100,000 and one in a million, depending on what survey you read,” Dr. Kleiner says. His advice: “Think of all the other things it could possibly be and rule those out before you say, yes, this is drug-induced liver injury. If it sounds like a big job, well, yes, it is. It’s probably the most challenging area in liver pathology.”
The Drug-Induced Liver Injury Network, or DILIN, was created in 2003 by the National Institute of Diabetes and Digestive and Kidney Diseases. It was in part “the result of a head-turning 2002 article [Lasser K, et al. JAMA. 287(17):2215–2220] that cited liver toxicity as the most common cause of a drug being withdrawn from the market or black boxed,” says Dr. Kleiner, who is a senior research physician in the Laboratory of Pathology at the National Cancer Institute, where he is also chief of the postmortem pathology section. “After that paper, there was a lot of attention by the FDA and Liver Diseases Research Branch of the NIDDK to see if they could explore this issue more.”
DILIN consists of one data coordinating center, at Duke University, and hepatologists working at a half-dozen clinical sites. There are prospective and retrospective studies; Dr. Kleiner works on the prospective side. “Patients suspected of having drug-induced liver injury and meeting certain criteria can be enrolled in the network’s study. They’re followed for up to two years with periodic evaluations. Clinicians collect biosamples, blood, DNA. And then when there is a liver biopsy—and if they can get re-cuts of the biopsy—it is sent, under code, to me.”
All of the drugs the network concerns itself with are considered to cause injuries that are idiosyncratic in nature. “These drugs are generally thought to be one-offs,” Dr. Kleiner says. “Even if patients have susceptibility to these, they may not develop injury. The incidence varies between about one in a thousand to one in a million patients taking the drugs. Because it’s such a low number, usually these drugs are not caught as potential problems during clinical trials.”
Read more...Whole Exome Sequencing Now Available in the NCI-COMPASS, Laboratory of Pathology
September 9, 2021
The NCI Laboratory of Pathology NCI-COMPASS Program recently has developed and validated tumor/normal clinical whole exome sequencing which performs whole exome analysis from tumor DNA and normal control DNA in one integrated workflow. During library preparation, enrichment chemistry is optimized to capture nucleic acid targets from formalin-fixed, paraffin-embedded (FFPE) tissue specimens. With the initial release, the molecular pathology report will include single nucleotide variants (SNVs), indels, tumor mutation burden (TMB) , copy number variants and eventually microsatellite instability (MSI) as well. A molecular pathology report will be completed for each order with the findings to clinical actionability based on AMP/ASCO/CAP/ACMG guidelines. Patient consent is required for this test to be performed.
While this test is primary designed for somatically acquired alterations in tumors, in the analysis of the normal DNA, we will report any clinically significant germline cancer predisposition variants within ~150 cancer predisposition genes, based on the current American College of Medical Genetics and Genomics (ACMG) guidelines will be reported separately from the somatic genomic findings. The germline variant test consent, pre-genetic test education, and post-genetic test counselling are available through the Clinical Cancer Genetics Program, Genetic Branch, CCR (contact to Kathleen Calzone, PhD, RN, AGN-BC, FAAN, 240-760-6178, calzonek@mail.nih.gov).
The test orders have been released in CRIS. Please refer CCR clinical SOP # ADGC-5 “Tumor/Normal Whole Exome Sequencing: Consenting, Ordering, and Obtaining Results” at https://ccrod.cancer.gov/confluence/pages/viewpage.action?pageId=73203825
Please contact Dr. Kenneth Aldape or Dr. Liqiang Xi for any questions regarding clinical issues.
Technical Specifications
|
Technical Information |
Whole Exome DNA and RNA NGS |
|
Sample Requirements |
Tumor specimen: FFPE block or 20 unstained slides with a minimum of 20% tumor cellularity Matched normal specimen: (blood or saliva) |
|
Tumor Enrichment (when necessary) |
Macrodissection to isolate and increase the number of tumor cells to improve variant detection sensitivity |
|
Number of Genes |
19,433 genes |
|
Capture Content |
45 Mb exonic content (≥98% of RefSeq, CCDS, and Ensembl coding regions |
|
Unique On-Target Reads |
88% |
|
Mean Target Coverage (DNA exome) |
140x for tumor and 59x for normal |
|
Variant Limit of Detection |
>95% for SNVs and Indels at ≥10% VAF |
|
Positive Percentage Agreement (PPA) with TSO500 Clinical Samples Pathogenic and Likely Pathogenic Variants Variant Unknown Significant |
98.8% for SNVs and 88.5% for indels at ≥10% VAF 94.4% for SNVs and 81.3% for indels at ≥10% VAF |
LP's Dr. David Kleiner and Dr. Stefania Pittaluga participate in NIH COVID-19 Lecture Series
August 25, 2020
COVID-19 Autopsy Findings: A joint effort between NYU Winthrop Hospital and NCI - What have we learned so far?
Since the outbreak of COVID-19 disease, caused by SARS-CoV-2 infection, many studies focusing on clinical course, outcome, clinical parameters, prognostic markers, treatment strategies have been published. Although most patients experience mild symptoms, some have serious complications—including diffuse alveolar damage, hemodynamic shock, acute kidney failure, cardiac injury, and arrhythmia—that contribute to the high mortality rate. Autopsies can offer a better understanding of the underlying pathophysiology. Unfortunately few autopsies were performed early in the pandemic because of the potential risks. Untreated patients who died of SARS-CoV-2 were rarely autopsied. Most of the published autopsy studies have focused on lung disease with a few describing findings in other organs. We will discuss our experience with the first set of patient autopsies performed at NYU Winthrop Hospital as well as some of our own experience here at the NIH. We will review some of the key findings in major organ systems accompanied by immunohistochemical and in situ hybridization studies that examined some of the cytokines/chemokines that have been implicated in the pathogenesis of this viral infection.
Videocast can be viewed here: https://videocast.nih.gov/watch=38104
David Kleiner, M.D., Ph.D., Senior Research Physician, Laboratory of Pathology, CCR, NCI, NIH
Stefania Pittaluga, M.D., Ph.D., Senior Research Physician, Laboratory of Pathology, CCR, NCI, NIH
LP Designated as Laboratory for NCI-MATCH
August 20, 2020
The Laboratory of Pathology participates with the NCI-MATCH Designated Laboratory. The NCI-MATCH (EAY131) phase 2 precision medicine clinical trial (NCT02465060) is evaluating the effectiveness of treatment that is directed by genomic profiling in patients with solid tumors, lymphomas or myelomas that have progressed following standard treatments expected to prolong survival, or for rare cancer types for which there is no standard treatment. Hopefully this will provide you and your patients more options or opportunities beyond your clinical studies at the CCR.
The Laboratory of Pathology, ECOG-ACRIN and the NCI-MATCH study team are collaborating to identify eligible patients for NCI-MATCH, based on genomic profiling results from the Laboratory of Pathology. Effective immediately, our TSO500 is replaced Oncomine Comprehensive Assay for screening for MATCH variants. If a variant(s) is found to be a qualifying tumor gene variant, the variant will be sent to the MATCH study team for further review and you as an NIH clinician will received a referral letter from the Laboratory of Pathology.
To learn more about NCI-MATCH, including clinical trial sites across the country, visit www.ecog-acrin.org/nci-match-eay131 and https://www.cancer.gov/about-cancer/treatment/clinical-trials/nci-supported/nci-match. Please contact Dr. Mark Raffeld, or Ms. Kayla O’Donnell (NCI-MATCH coordinator) if there are any questions.
This is an update to an original article posted October 30, 2019.
CC Grand Rounds John Laws Decker Lecture -
David E. Kleiner, MD, PhD - Distinguished Clinical Teacher Awardee 2024 Award
You are cordially invited to this reception, to celebrate Dr. David E. Kleiner as the 2024 Distinguished Clinical Teacher Awardee
When: Wednesday June 12th at 1-2pm
Where: Medical Board Room
To RSVP please sign up here by Friday May 31
The reception is after his 12pm CC Grand Rounds Decker – DCTA Lecture in Lipsett
Steatotic Liver Disease: From First Description to First Drug Approval Through The Eyes Of The Pathologist
If you have questions, contact Rita Stevens at rita.stevens@nih.gov
Elaine S. Jaffe, M.D. Named NIH Distinguished Investigator
October 29, 2019
Congratulations Dr. Jaffe!
TruSight Oncology ~500 Gene Panel (TSO500) Is Now Available
Kenneth Aldape, MD
October 17, 2019
I am pleased to announce a major step forward in the NCI’s Comprehensive Oncologic Molecular Pathology and Sequencing Service (NCI-COMPASS) of the Laboratory of Pathology.
Effective immediately, a new clinical molecular test, TruSight Oncology ~500 Gene Panel (TSO500) is now available on the Molecular Pathology CRIS order menu, which is relevant for a large variety of solid tumors.
The TSO500 is our next-generation sequencing (NGS) assay that analyzes cancer-relevant genes from both DNA and RNA in one integrated workflow. During library preparation, enrichment chemistry is optimized to capture nucleic acid targets from formalin-fixed, paraffin-embedded (FFPE) tissues. With simultaneous analysis of both DNA and RNA, various types of biomarkers relevant to a given tumor type (single nucleotide variants (SNVs), indels, fusions, splice variants, tumor mutation burden (TMB), and microsatellite instability (MSI)) can be assessed from the same sample in a single assay. The RNA panel uses a probe design that enables capture of both known fusions and novel fusion partners. The TSO500 panel includes 523 genes for DNA mutation detection and 55 genes for fusion and splice variant detection.
The new molecular pathology report is also incorporated with reporting software for clinical actionability as Tier levels of FDA-approved drug and clinical trials, and pathogenicity based on AMP/ASCO/CAP/ACMG guidelines.
NIH staff may access the CRIS order instructions and full gene list of TSO500 on the NCI-COMPASS webpage. Although Oncomine Assay is still available in CRIS menu for a period, for most applications, the TSO500 can replace the Oncomine Assay for all tumor types, given that it has added features including TMB score, and MSI. In the near future, copy number variation (CNV) will be available. There will be no need to order the Oncomine assay in the future if the TSO500 panel is requested.
Please contact Dr. Liqiang Xi if there are any questions regarding this assay, or Ms. Kayla O’Donnell for question about ordering Molecular Assays.
NCI-Connections features LP's Dr. Kenneth Aldape
October 10, 2019
Why a Precise Diagnosis is Vital to Treat Brain and Spine Tumors
Most people first learn they have a tumor from their primary care doctor. They are then referred to neuro-oncologists or doctors who have expertise in their tumor type. Part of this specialized healthcare team is a neuropathologist, a doctor who examines brain and spine tumor tissue to make a precise diagnosis.
“We come up with the best definition of the tumor possible based on the specific changes we see in the tumor under the microscope,” says Kenneth Aldape, M.D., Ph.D., senior investigator and chief of the Laboratory of Pathology in the Center for Cancer Research at NCI.
Dr. Aldape studies the genomic and epigenomic alterations in brain and spine tumors – or the way genes behave and change in cancer. He characterizes the biology of specific genomic alterations and how they contribute to the development of the disease. He also studies how they impact treatment resistance of aggressive brain tumors.
The goal of understanding these alterations is to be able to identify and better classify tumor types. “We believe that classifying brain tumors based on their biology will lead to a greater understanding of why specific tumor subtypes may be more or less sensitive to therapies,” Dr. Aldape says.
He is also pushing for precision diagnosis for patients because of the rate of diagnostic discrepancy. “We want to reduce errors because without a proper diagnosis a lot of precision medicine may not go as well as planned,” Dr. Aldape says.
Making an Accurate Diagnosis
Dr. Aldape works on the NCI-CONNECT healthcare team to help accurately diagnose rare brain and spine tumors. “Some of these rare tumors can be challenging to diagnose because they are so rare,” Dr. Aldape says.
At NIH, all patients who visit the Neuro-Oncology Branch have their diagnosis reviewed by a neuropathologist. Dr. Aldape receives tumor tissue in the form of a tissue block or unstained slides from the hospital where a patient had his or her surgery. If a patient has not had surgery or a biopsy, he or she can consult with NIH’s neurosurgical oncology team. Dr. Aldape examines the tissue closely under a microscope looking at alterations in the tissue. Read more...
Laboratory of Pathology's June 2019 Newsletter
Kenneth Aldape, MD
June 5, 2019
It is axiomatic that precision medicine in oncology will require advances in precision diagnostics, and advances in our understanding of tumor genomics provide a promising approach towards improvements in precision cancer diagnostics. Individuals bearing tumors with specific molecular alterations, such as gene mutations, amplifications and translocations, and microsatellite instability can be identified using molecular diagnostic tools available in pathology departments, and recent clinical trials for molecularly targeted agents and immunotherapies have been shown to produce unprecedented extended survival in patients with molecularly-defined tumors. While it remains to be determined whether establishing a comprehensive tumor molecular profile of the tumor improves outcome following therapy overall, the evidence to date indicates that this approach represents a promising and dynamic pathway to this goal for specific cancer subsets, and with new data and trials that emerge on an almost a weekly basis, the future remains bright in this area.
Read more...Flow Cytometry Section of the Laboratory of Pathology, NCI Receives an Award from Wiley Publishing
June 3, 2019
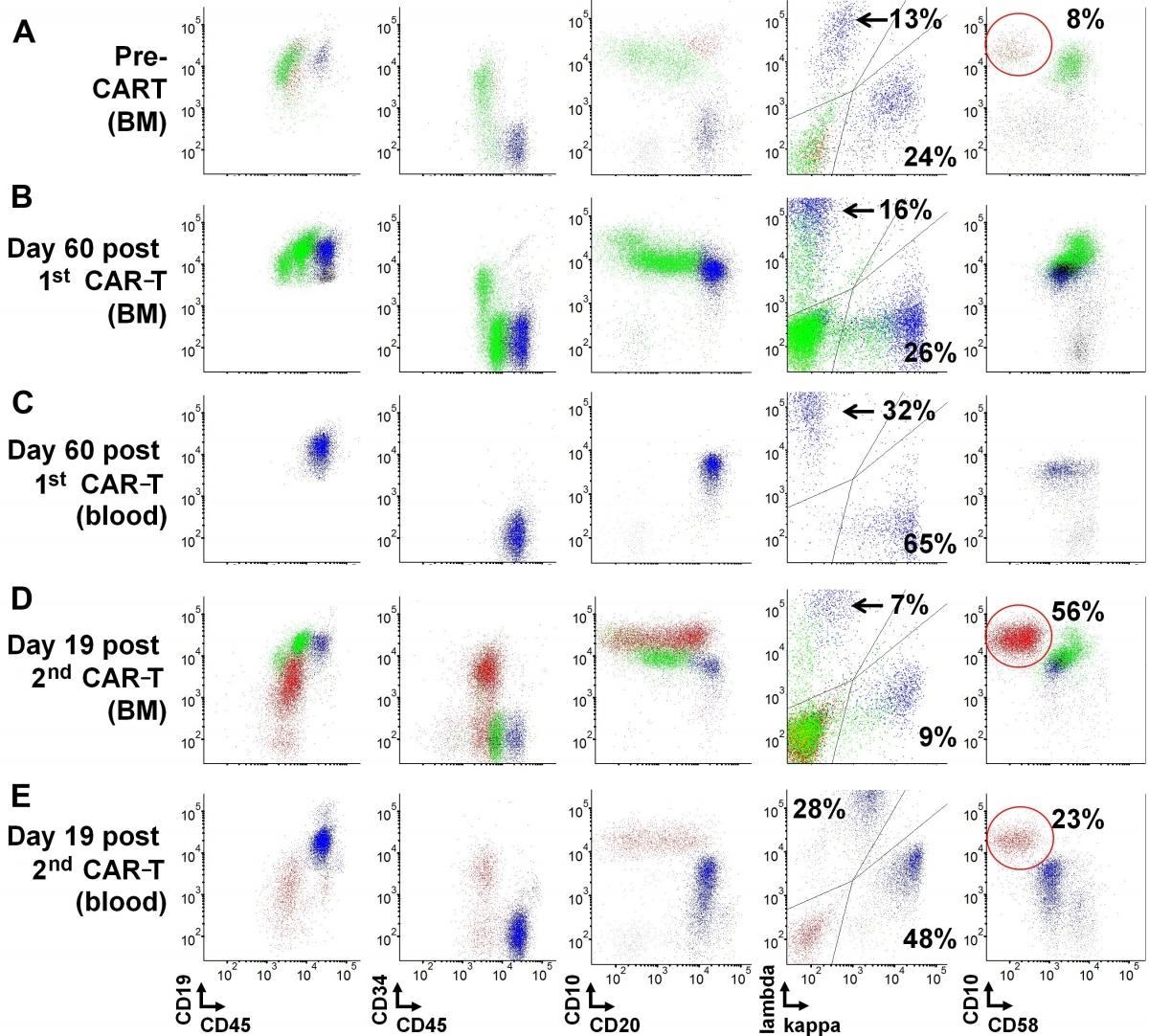
Image: Marrow recovery and circulating transitional B-cells in a lymphoblastic leukemia patient post CAR-T cell therapy.
The NCI LP Flow Cytometry section achieved recognition for a Top 20 Article in Cytometry Part B: Clinical Cytometry for their original work entitled, “Early recovery of circulating immature B cells in B-lymphoblastic leukemia patients after CD19 targeted CAR T cell therapy: A pitfall for minimal residual disease detection”. This publication was recognized as one of the journal's top downloaded recent papers. Amongst articles published between January 2017 and December 2018, this work received some of the most downloads in the 12 months following its online publication. This original work highlighted an unusual B-cell marrow recovery response resulting in a notable population of circulating immature B-lymphocytes. These immature B-lymphocytes, if not carefully evaluated, might be mistaken for residual B-lymphoblastic leukemia, due to their immunophenotypic features. In fact, the presence of these cells, likely representing transitional B-cells, may reflect brisk regeneration of the B-cell compartment and bone marrow recovery following treatment with CAR T-cell therapy. Furthermore, the work details specific clinical flow cytometric strategies that can be used for evaluation and prevent a mis-diagnosis of B-lymphoblastic leukemia. According to Wiley publishing, the work generated immediate impact and visibility, contributing significantly to the advancement of the field of clinical flow cytometry.
Methylation Classifier Now in Use as a Clinical Diagnostic Tool
May 24, 2019
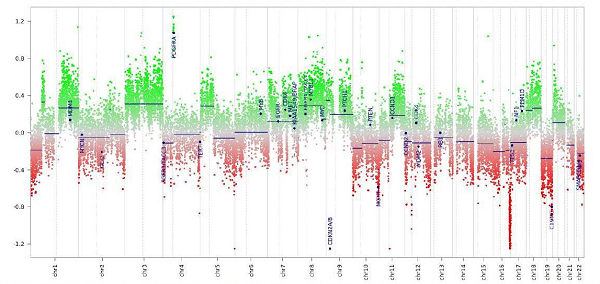
Sample image of a patient’s results produced by the tumor methylation diagnostic tool.
The NCI Laboratory of Pathology has recently begun to use a new clinically-reportable diagnostic tool that uses genome-wide DNA methylation profiling as a diagnostic for tumors of the central nervous system. The validated tool is based, in part, on data published in a recent Nature study that showed tumor methylation profiles can provide definitive evidence to complement and refine morphology-based diagnostics in tumors of the brain and spinal cord. In the study, methylation data resulted in a change in diagnosis for 129 cases (12%) of the cohort.
The NCI Laboratory of Pathology is poised to become a diagnostic reference center to implement this tool for diagnostically challenging neuropathology cases. Going forward, it is likely that new methylation-based classifiers will emerge for additional tumor types and we are poised to lead in this area. Areas of future growth include the implementation of clinical whole-exome sequencing, RNAseq gene expression diagnostics, and a dynamic liquid biopsy program.
For more information about access to the Laboratory of Pathology's methylation diagnostic tool, please email the Methylation Profiling Group.
Elaine S. Jaffe, M.D., Receives the 2019 USCAP Board's Distinguished Pathologist Award
April 11, 2019
Elaine Jaffe, M.D., Senior Investigator in the Laboratory of Pathology, has received the 2019 United States and Canadian Academy of Pathology (USCAP) Board’s Distinguished Pathologist award. The award recognizes an individual for making major contributions to pathology over the years.
The Board’s Distinguished Pathologist award was established by the Board of Directors of USCAP for recognition of distinguished service in the development of the discipline of pathology. This award is presented to an individual who is recognized for making major contributions to pathology over the years. Read more...
LP Chief Ken Aldape Participates in NCI Facebook Live Event
"Genomics: How DNA Can Inform Cancer Diagnosis and Treatment"
February 14, 2019
The National Cancer Institute hosted a Facebook Live on February 14, at 3 p.m. ET. The event featured NCI subject matter experts Kenneth Aldape, M.D., of the Center for Cancer Research, Megan Frone, M.S., C.G.C., of the Division of Cancer Epidemiology and Genetics, and Lyndsay Harris, M.D., of the Cancer Diagnosis Program. They discussed the role of genetics in cancer diagnosis and treatment. At the end of the event, experts answered live questions from viewers.

Research
We support the research mission of the NCI and the NIH by investigating the biology and genetics of cancer and other diseases, developing and applying leading edge technology to diagnostic pathology, and providing collaborative support for clinical research protocols.
Alumni
Training
Pathology Training and Fellowship Programs
Through the training of residents and fellows in diagnostics and research, LP's Training Programs create future leaders in clinical and experimental pathology. As an organization, we share our expertise and teaching with the clinical and scientific community.
Anatomic Pathology Residency Training Program
The Laboratory of Pathology (LP) offers a multifaceted ACGME-accredited residency training program in Anatomic Pathology at the NIH Clinical Center, the site of intramural clinical research for the NIH, and home to more than 1500 clinical research protocols. Excelling in both clinical diagnosis and translational research, the department provides a stimulating intellectual environment for the resident interested in an academic career. Learn more...
Cytopathology Fellowship
The LP's Cytopathology Section provides diagnostic cytopathology services to the NIH and its associated clinical services in the NIH Clinical Center. The relatively high frequency of pathologic findings combined with the diversity of types of exfoliative and fine needle aspiration (FNA) specimens seen in our Section provide a broad experience in diagnostic cytopathology. The goals of this fellowship are to develop a strong foundation in diagnostic cytopathology and introduce clinically oriented physicians to current research techniques. Learn more...
Hematopathology Fellowship
The LP's Hematopathology Section offers a fully ACGME-accredited fellowship in hematopathology, which provides broad exposure to the diagnostic and investigative aspects of neoplastic hematopathology. Material is derived principally from an active in-house treatment program for both adult and pediatric hematologic malignancies. In addition, approximately over 2000 challenging cases are submitted in consultation each year. Learn more...
Pathology Research Fellowships
Pathology research fellowship positions are available in the Laboratory for physicians who have recently completed a pathology residency program. This fellowship is intended to give pathologists sustained and focused training in an investigative area in basic, translational or clinical science in preparation for a career in academic pathology. Learn more...
Cancer Genomics Pathology Research Fellowships
Research fellowship positions in the area of cancer genomics pathology are available in the Laboratory for physicians who have recently completed a pathology residency program. This fellowship is intended to give pathologists sustained and focused training in cancer genomics in preparation for a career in academic pathology. Learn more...