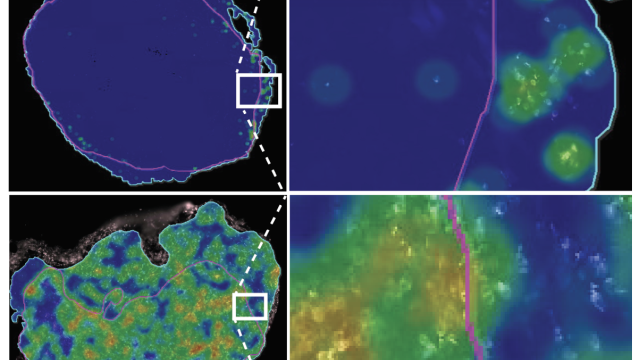
An innovative strategy for mining cancer genomes finds enzymes with tumor-suppressing roles.
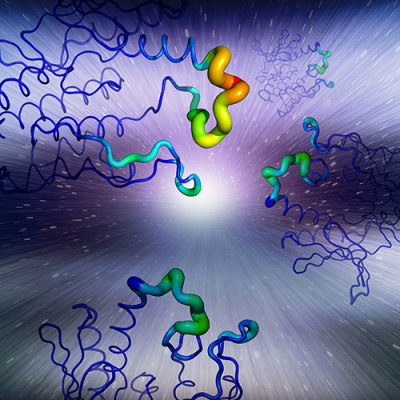
Advanced molecular dynamics is a method that helps researchers study the physical movements of molecules. The technique helps scientists to predict if cancer mutations will alter the structure of an enzyme, such as a kinase, in a manner that may alter the enzyme’s function to promote cancer. These two different structures compare the normal enzyme (top) and a cancer-mutant enzyme (G265D) for the kinase MAP2K4 (bottom). The cancer mutant (G265D) has dramatically less movement, indicating the enzyme will not be functional, which is consistent with mutations in tumor-suppressing enzymes.
Credit: John Brognard, CCR, NCI, NIH
Cancer biologists have long been interested in a family of enzymes called kinases. More than 500 different kinases are found in human cells where they transmit critical signals by adding phosphate groups to proteins. A handful of kinases have received a great deal of attention in cancer research because they are disrupted by genetic mutations that are well known to drive cancer. Most, however, remain poorly characterized. Now, CCR scientists have devised a clever new way to identify additional cancer-relevant members of this prominent class of proteins.
John Brognard, Ph.D., Investigator in CCR’s Laboratory of Cell and Developmental Signaling, and Andrew Hudson, M.D., a former clinical fellow in his lab, devised an innovative strategy to sift through data describing the tens of thousands of mutations that have been found in cancer cells. Using data from The Cancer Genome Atlas, which was supported by NIH, and the Cancer Cell Line Encyclopedia, developed at the Broad Institute, their team searched for mutations in kinase genes in more than 11,000 patient tumors and 1,000 cancer cell lines.
Because they sought mutations that impact kinase function, Brognard’s team limited its search to stretches of sequence known to be critical for the enzymes’ activities, filtering out all other mutations. In this way, they identified several kinases with high frequencies of functional mutations across the large set of samples.
From this group, they focused on a kinase called MAP2K7, a component of the JNK signaling pathway that is known to be capable of both spurring cancerous growth and reining it in. The team’s analysis showed that functional mutations in MAP2K7 were common among tumor samples from patients with gastric cancer. They went on to demonstrate that intact JNK signaling is important for keeping growth of gastric cells in check.
The impact of this work, which the researchers reported in Science Signaling, reaches beyond the important leads it has already turned up. The data-mining approach is not limited to kinases, Brognard says, and can now be used to search existing genomic data for cancer-driving mutations in other families of enzymes.



