Tumor cells rely on genes to escape immune control. Understanding the genes tumor cells rely on to escape immune control could guide the way to immunotherapies that work for more patients.
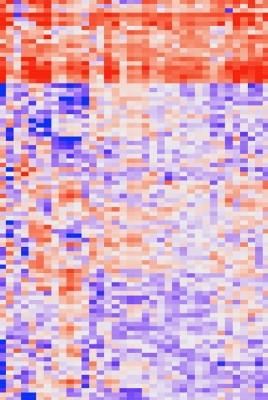
This heat map graph shows the gene expression profiles of over 11,000 different tumors. The cluster of genes shown in red correlates with the presence of T cells.
Credit: Nicholas Restifo, CCR, NCI, NIH
Immunotherapies are changing the landscape of cancer treatment. They work by empowering a patient’s immune system to attack difficult-to-treat cancers, often leading to complete disappearance of tumors. But many patients still fail to respond to these innovative treatments, and developing immunotherapies that work for more people is a high priority.
Senior Investigator Nicholas Restifo, M.D., from CCR’s Surgery Branch, and his team of researchers have addressed this issue in a systematic fashion and have created a complete compendium of human genes that affect how well immunotherapy works.
Restifo and his colleagues tested all of the nearly 25,000 protein-coding genes in the human genome to determine which ones must be working in tumor cells to make them vulnerable to immune attack. In a landmark publication in Nature, the team reported on 19 genes that appear to be essential for a successful anticancer immune response in patients. Their findings suggest that tumors that accumulate mutations in any of these genes might become less susceptible to immune control.
Current immunotherapies are designed to boost the ability of the immune system’s T cells to find and destroy tumors. Restifo and colleagues used CRISPR, a powerful gene-editing technology, to generate melanoma cells with individual genes switched off. Then they allowed T cells to mingle with the modified melanoma cells and observed how the immune cells performed when their targets lacked any given gene. In this way, the researchers identified more than 100 genes that appeared to play a role in the T-cell response.
The team then accessed data from The Cancer Genome Atlas (TCGA), a publicly available resource collaboratively managed by NCI and the National Human Genome Research Institute. Using DNA sequence information from more than 11,000 patient tumors in TCGA, the team looked for evidence that the genes they identified in their laboratory experiments played a similar role in patients. By mining that data for patterns of gene activity that typically accompany cell death, the researchers linked 19 of the genes on their list to tumors’ ability to avoid immune attack in patients.
Restifo’s team has already zeroed in on one of these genes, APLNR, which encodes a cellular signaling protein, and discovered that it is critical for processing the markers on the surface of tumor cells that alert T cells to their identity. They found APLNR mutations in tumors from patients whose cancers had not responded to immunotherapy and showed that when melanoma cells contain mutations in APLNR, the immune system has a hard time finding tumors and keeping their growth in check.
Further studies should begin to unravel the other genes’ contributions to a tumor’s susceptibility to immune control. In addition to shedding light on how immunotherapy works, in the near-term, the catalogue of genes identified will help researchers predict which patients are most likely to benefit from specific immunotherapies. By exploiting the tumors’ specific strategies for escaping the immune system, the catalogue could point the way to new therapies designed to counter those tactics.
Reference
Patel SJ, et al. Nature. 2017 Aug 31;548(7669):537-542.


