Eugene Valkov, D.Phil.
- Center for Cancer Research
- National Cancer Institute
- Building 560, Room 11-83C
- Frederick, MD 21702
- +1 (301) 846-1823
- eugene.valkov@nih.gov
RESEARCH SUMMARY
What determines the fate of an mRNA at the end of the gene expression pathway? Our laboratory addresses this question by dissecting the molecular mechanisms that govern mRNA stability and decay. We combine biochemistry, structural biology, and computational approaches to build detailed, mechanistic models of how multiprotein complexes regulate transcript lifetimes.
We have established robust platforms for the recombinant production and characterization of human RNA regulatory factors and complexes, enabling systematic reconstitution of the mRNA decay machinery. These studies integrate high-resolution cryo-electron microscopy, machine learning-based computational tools, and rigorous biochemical and biophysical validation of protein-RNA interfaces, affinities, stoichiometries, and kinetics.
Although our work is rooted in understanding fundamental principles of post-transcriptional control that are shared across all eukaryotes, we are increasingly applying our expertise to contexts where gene expression becomes dysregulated, including cancer and chronic disease. By studying how RNA decay is rewired in these states, we aim to uncover novel therapeutic targets and lay the groundwork for innovative treatment strategies.
Areas of Expertise

Eugene Valkov, D.Phil.
Research
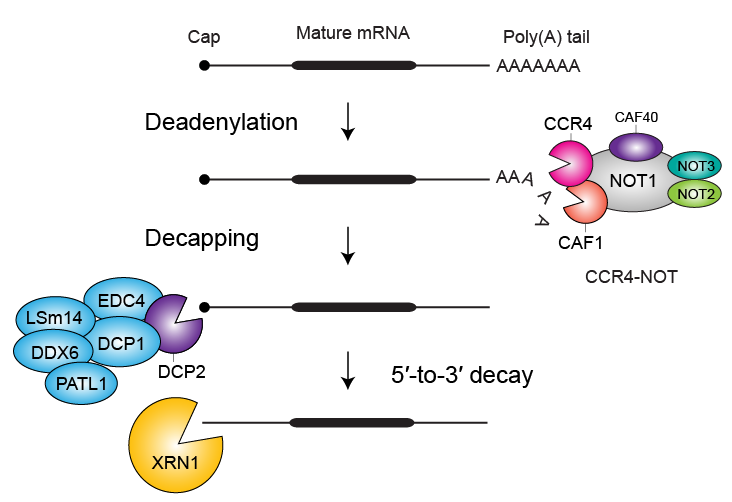
Gene expression is commonly portrayed as a linear flow from chromatin to ribosomes. Yet, it is the balance between the synthesis and decay of messenger RNA (mRNA) that ultimately determines protein output. The degradation arm of this balance, particularly the stepwise shortening of the poly(A) tail (deadenylation) that precedes mRNA destruction, is emerging as a decisive checkpoint in cell-state transitions, metabolic adaptation, and oncogenic re-programming. Despite its importance, mRNA decay has lagged behind transcription and translation in mechanistic depth, primarily because the unstable, heterogeneous ribonuclease assemblies that initiate decay have resisted biochemical reconstitution.
The last decade has seen a paradigm change. Structural snapshots, single-nucleotide kinetic profiling, and the first complete reconstitutions of the human mRNA decay complexes now provide a solid mechanistic foundation for studies in how cells decide whether a transcript will be translated, stored, or eliminated. My laboratory, initially located at the Max Planck Institute and now at the National Cancer Institute since 2019, has been at the forefront of this shift. We integrate structural biochemistry, systems-level genomics, chemical biology, and advanced protein design to understand and ultimately manipulate the molecular logic that connects poly(A) tail length, RNA-binding proteins (RBPs), and decay factor recruitment to control gene expression programs in development and cancer.
Our research centers on a fundamental question: How do dynamic, modular interactions within the CCR4–NOT complex and with other factors dictate the fate of cellular mRNAs, and how are these processes rewired in development and disease?
Looking ahead, our long-term goal is to determine whether natural or engineered modulators of activity, as well as RNA-binding proteins that recruit CCR4-NOT, can be harnessed to therapeutically reshape transcriptomes in cancer and chronic disease.
Our overarching hypothesis is that low-affinity, multivalent, and highly dynamic interfaces, rather than single high-affinity contacts, provide the speed, reversibility, and context adaptability essential for post-transcriptional control. By mapping, quantifying, and re-engineering these interfaces, we aim to uncover fundamental principles of RNA regulation and identify novel drug targets with the potential to seed first-in-class therapeutic strategies.
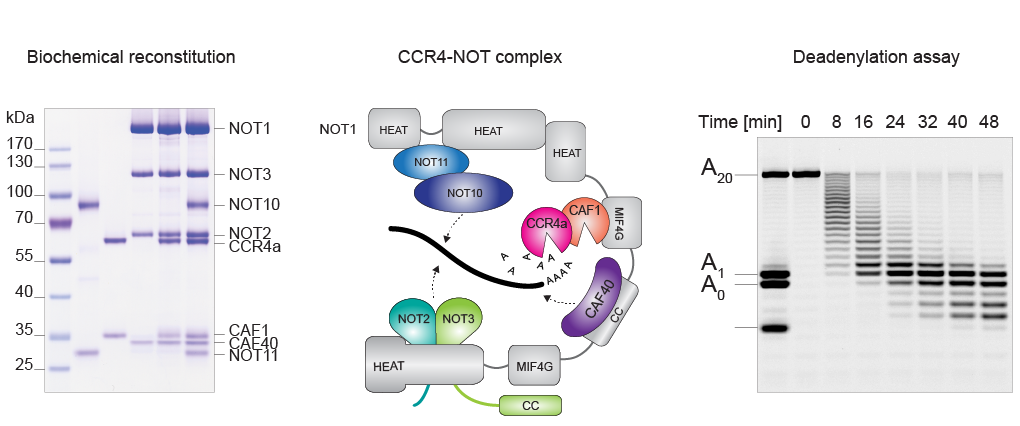
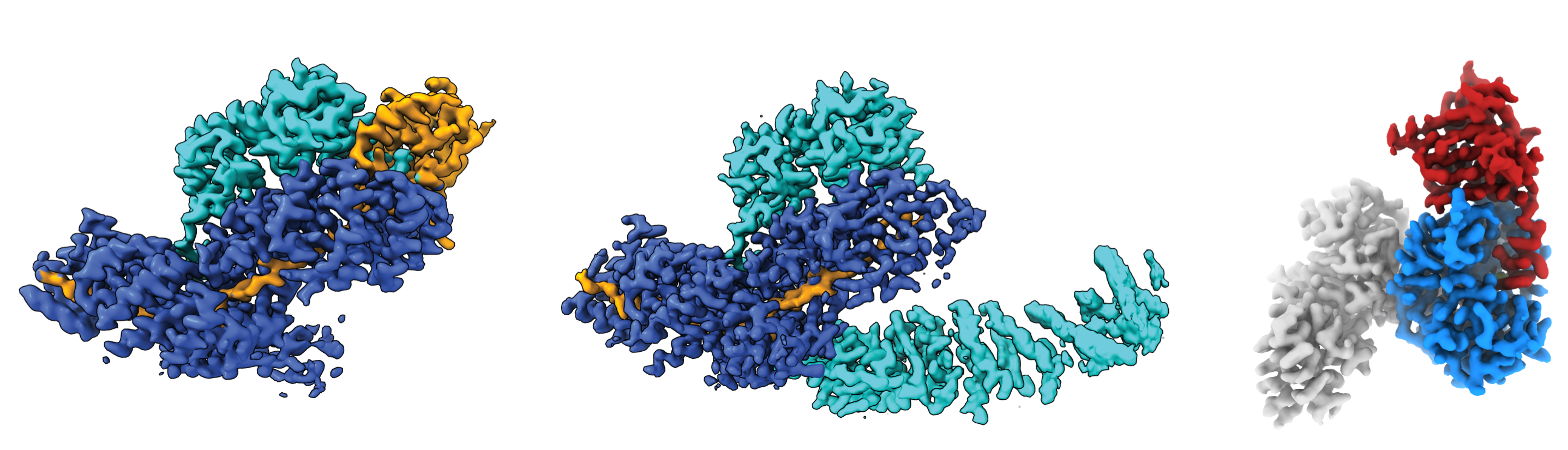
Our primary approach to studying these molecular events involves reconstituting the relevant regulatory macromolecular complexes from highly purified recombinant components and examining them in vitro. To this end, we employ biochemical assays and biophysical approaches to study the interactions between the components using the instrumentation and facilities available at the Biophysical Resource, part of the Center for Structural Biology. To gain a deeper understanding of the molecular mechanism, we determine high-resolution cryo-EM structures through single-particle analysis using data collected on the Titan Krios G4 and Talos Arctica microscopes available on our campus. Among our recent achievements is the reconstitution of the complete human CCR4-NOT deadenylation complex and the entire network of principal human decapping factors. These unique biochemical tools and reagents enable us to study the 5′-to-3′ mRNA decay pathway with unprecedented precision and control using structural, biochemical, and cellular approaches.
To relate biochemical and structural insights to function, we test our derived hypotheses directly in human cells, for example, by complementing different variants in cell lines where the expression of individual molecular factors has been knocked out by CRISPR gene editing. This approach has proved very powerful in validating and mapping interactions in a cellular context. We utilize high-throughput short-read sequencing, combined with bioinformatic analysis, to survey the dynamics of the cellular transcriptome and translatome. We also employ long-read direct RNA sequencing approaches to study mRNA metabolism using Oxford Nanopore instruments available at the RNA Biology Laboratory.
Relevance to Cancer
Dysregulation of post-transcriptional control is a hallmark of numerous human diseases, including autoimmunity, neurodegeneration, and, most prominently, cancer. By uncovering the fundamental mechanisms that govern mRNA turnover and translation, our research provides critical insights into how these processes are perturbed during oncogenic transformation and in age-related chronic diseases. A deeper mechanistic understanding of RNA regulation not only advances basic biology but also opens new therapeutic avenues, offering fresh strategies to intervene in tumor initiation, progression, and metastasis.
Publications
Biography

Eugene Valkov, D.Phil.
Eugene Valkov received his doctorate in molecular biophysics from the University of Oxford. As a postdoctoral researcher at the MRC Laboratory of Molecular Biology in Cambridge, he investigated the molecular mechanisms of mRNA export and decay. In 2014, he joined the Max Planck Institute in Tübingen as a staff scientist with Elisa Izaurralde and was promoted to group leader in 2017. He joined the RNA Biology Laboratory in 2019.
Job Vacancies
We have no open positions in our group at this time, please check back later.
To see all available positions at CCR, take a look at our Careers page. You can also subscribe to receive CCR's latest job and training opportunities in your inbox.
Team
News
August, 2025. In a Nature Communications study, led by Filip, we demonstrate that tristetraprolin (TTP) employs multivalent, phosphorylation-regulated interactions with the CCR4-NOT deadenylase and PABPC1 to orchestrate rapid, AU-rich element-specific mRNA deadenylation and decay. Notably, Josh, a former postbac, and Keiko, a former summer intern, were both recognized for their contributions through co-authorship. Carried out in collaboration with Perry Blackshear at the NIEHS.
August, 2025. Claire and Eugene contributed to a study led by Hans Elmlund and our colleagues in the Center for Structural Biology, published in Acta Crystallographica Section D, which presents a probabilistic, real-time capable cryo-EM ab initio reconstruction method (included in the SIMPLE software suite) that simultaneously determines particle orientations, partitions heterogeneous states, and applies adaptive non-uniform regularization to enhance accuracy and computational efficiency.
July, 2025. In a real team effort, using cryo-EM, biochemistry, and cellular analyses of mRNA poly(A) tails by Nanopore sequencing, we describe in a Nature Communications paper how that the Legionella effector PieF inhibits host mRNA decay by binding the NOT7 and NOT8 subunits of the CCR4-NOT deadenylase by blocking assembly of the active complex and arresting mRNA deadenylation to disrupt cell cycle progression in human cells. Yevgen, David, and Claire are all co-first authors, in addition to Justin Deme from Susan Lea's group at the Center for Structural Biology. Filip also contributed, as well as Anna Valkov and Sergey Tarasov, with their expertise.
July, 2025. Yevgen and Eugene contributed to a collaborative study by Quaid Morris and colleagues, published in Nature Biotechnology, which introduces a comprehensive, freely accessible resource of RNA-binding protein motifs encompassing 34,746 RBPs across 690 eukaryotic species. This was generated using the novel Joint Protein-Ligand Embedding (JPLE) algorithm, which has vastly expanded motif coverage, illuminated evolutionary dynamics, and enhanced our ability to infer post-transcriptional regulatory functions.
July, 2025. Annie, Claire, and Jade all finished their postbac time with us and moved on to their next career stage.
June, 2025. In a collaborative study with Russell Vance and colleagues, published in Nature, we contributed to the discovery that the transcriptional repressor SP140 controls antiviral immunity not by inhibiting IFNB1 transcription, but by suppressing the expression of a previously uncharacterized stabilizer called RESIST, which otherwise protects IFNB1 mRNA from degradation by TTP and the CCR4-NOT complex. David and Filip contributed to this work by characterizing RESIST interactions with the CCR4-NOT and exploring how this may compete with TTP.
May, 2025. Claire accepted an offer to join the MD/PhD program at UCSF, and Annie chose to join the PhD program at the University of Chicago. Congrats, Claire and Annie!
December, 2024. Eric left the lab to explore careers in the biotech industry after completing his postdoctoral fellowship.
October, 2024. Eugene gave a talk on PieF/CCR4-NOT mechanism at the EMBO | EMBL Symposium: The Complex Life of RNA at Heidelberg, Germany.
October, 2024. Eugene gave a seminar at the Memorial Sloan Kettering Cancer Center in New York City. Thank you to Chris Lima for the invitation and excellent hosting!
September, 2024. The NCI Site Visit panel reviewed our research program, together with our colleagues at the RNA Biology Laboratory. The reviewers were very enthusiastic about our research objectives and recommended that we continue!
August, 2024. Eugene and Filip both gave talks and Eugene chaired a session at The Post-transcriptional Control of Gene Expression Conference on RNA Decay in Lisbon, Portugal. Eugene presented the mechanism of the PieF-mediated inhibition of CCR4-NOT, while Filip described his work on TTP-mediated repression.
August, 2024. Our summer intern, Jennifer, left the lab to continue her undergraduate studies at the University of Maryland.
July, 2024. David and Yevgen both won a Fellows Award for Research Excellence (FARE), which provides supplementary funding for them to attend a conference of their choice to present their work. Congrats!
May, 2024. Jennifer Owen, an undergraduate student pursuing a Neuroscience Major on the Pre-Med Track at the University of Maryland, joined the lab as part of the Summer Internship Program to get some research experience. Welcome, Jennifer!
May, 2024. Eugene gave a seminar at the Department of Biochemistry at McGill University in Montreal, Canada. Thank you to Marc Fabian and Will Pastor for the invitation and fantastic hosting!
April, 2024. Together with Alicia Bicknell and colleagues at Moderna, we show that high ribosome loads result in less overall protein output from therapeutic mRNAs due to increased translation-dependent mRNA decay. The implication is that modulating the translation initiation rate to achieve moderate ribosome loads and longer mRNA lifetimes can improve the efficacy of therapeutic mRNAs. Yevgen tested the 3' stabilizing mRNA modification in our in vitro deadenylation system. Out now in Cell Reports.
April, 2024. Our collaborative work with Jernej Murn and other colleagues is now out in Nature Communications. We investigated the molecular mechanism of an RNA-binding protein, Unkempt, and its interactions with CCR4-NOT and PABPC1 in regulating translation and cell polarization. From our lab, David characterized Unkempt interactions in vitro; Josh established the purification of full-length Unkempt and carried preliminary assays; Khadija performed some of the mRNA poly(A) tail analysis using Nanopore.
February, 2024. Our study on deadenylation kinetics, done in collaboration with Narry Kim's group, was published in Nature Structural & Molecular Biology. We quantitated the effect of the incorporation of non-adenosines in mRNA poly(A) tails, or 'mixed tailing,' which has been observed in vertebrates and viruses, by developing an analytical framework to monitor deadenylation single-nucleotide resolution using an in vitro deadenylation system reconstituted with the complete human CCR4-NOT complex. This study reveals the molecular basis of tail sequence-dependent regulation of mRNA stability, which has implications for improving mRNA-based vaccines and therapeutics. Yevgen performed all the experimental work in this study.
February, 2024. Tanner Myers joined us as a postdoc after completing his doctoral work on bacterial ribonucleases with Wade Winkler at the University of Maryland. Welcome, Tanner!
December, 2023. Eugene gave a seminar at the SickKids Hospital in Toronto. Thank you to Julie Forman-Kay for the invitation and the opportunity to learn about all the great work done at SickKids.
October, 2023. Deadenylation: Methods and Protocols, co-edited by Eugene and Aaron Goldstrohm. Yevgen and Filip contributed a chapter each. Many thanks to all the leading laboratories in the field for contributing their latest methods and expertise!
September, 2023. Filip was awarded a postdoctoral fellowship from the German Research Foundation (DFG) as part of the Walter Benjamin program. Many thanks to the reviewers for their very kind comments, including "I cannot imagine a better place to carry out the proposed project" and "I rate this proposal among the top 5% and recommend funding with highest priority for a period of 24 months.“ Congrats, Filip!
August, 2023. Our summer intern, Keiko, left the lab to continue her undergraduate studies at the University of Maryland. Good luck, Keiko, with your studies, and we hope to see you again soon!
August, 2023. Annie Corbett and Claire Piczak joined the lab as postbacs. Both will work on biochemical and structural studies of RNA-binding proteins. Welcome, Annie and Claire!
August, 2023. The NIGMS funded our collaborative R01 grant with Aaron Goldstrohm on mRNA regulation by human Pumilio proteins. The NCI/CCR provided supplementary funding for our requested resources via a unique mechanism that seeks to promote intramural/extramural collaboration.
July, 2023. Cryo-EM structures of human and chicken NOT1:10:11 ternary subcomplexes of CCR4-NOT are published in Communications Biology. Thanks to all, especially our collaborators Susan Lea and Justin Deme, for introducing us to cryo-EM! Yevgen and Filip prepared the samples.
July, 2023. Jade Witter joined the lab as a postbac and part of the NCI Intramural Continuing Umbrella of Research Experiences (iCURE) program. Jade will work on a collaborative project with Jay Schneekloth's group at the Chemical Biology Laboratory. Welcome, Jade!
July, 2023. Wataru left the lab to join Daniel Kiss as a staff scientist at Houston Methodist Research Institute. Thanks, Wataru, for your hard work and contributions, and we wish you the best in your future endeavors.
June, 2023. Keiko Luke, an undergraduate student in biological sciences at the University of Maryland, joined the lab as part of the Summer Internship Program to get some research experience. Welcome, Keiko!
May, 2023. Filip's study of RBP-directed deadenylation is published in Nucleic Acids Research — part of this work Filip carried out with us at the NCI. Congrats, Filip, on your first first-author publication!
April, 2023. Leah accepted an offer to join Jan Lowe's lab as part of the International PhD Program at the MRC Laboratory of Molecular Biology in Cambridge, U.K. Congrats, Leah!
March, 2023. Eugene gave a seminar at University of California at Riverside. Thank you to Ted Karginov, Jernej Murn, Sihem Cheloufi, and all the staff and students at UC Riverside for a wonderful visit and for sharing their science!
March, 2023. Yevgen was among two students recognized this year with the Center for Cancer Research Outstanding PhD Student Award for his stellar doctoral work on the biochemistry of the human CCR4-NOT complex. Congratulations, Yevgen!
March, 2023. Khadija left the lab to return to her family in Morocco. Thanks, Khadija, for all your contributions to several projects, and we wish you the best in your future endeavors.
December, 2022. Adrianna left the lab to pursue a medical degree at the University of Queensland, Australia. Best of luck with your medical studies, Adrianna!
December, 2022. Yevgen defended his Ph.D. thesis virtually at the University of Tübingen, Germany, with an overall mark of 1.0 and a distinction of Magna Cum Laude. Many thanks to the chair of the committee, Professor Dirk Schwarzer, and the members of Yevgen's thesis committee, Professors Thilo Stehle and Ralf-Peter Jansen, for their time and support throughout the years.
December, 2022. Eugene gave an invited lecture at Ohio State University in Columbus, Ohio. Many thanks to Mike Kearse for the kind invitation and to all the staff and students at Ohio State for a great visit!
December, 2022. The Current Opinion in Structural Biology collection on Protein Nucleic Acid Interactions (2022), where Eugene served as a Guest Co-Editor, is published.
October, 2022. Eugene gave an invited lecture at Moderna headquarters in Boston, Massachusetts. Many thanks to Alicia Bicknell and her colleagues for a kind invitation and a great visit. We look to working together to help develop future mRNA-based therapeutics!
July, 2022. Adrianna obtained a Cancer Research Training Award (CRTA) from the NCI Intramural Continuing Umbrella of Research Experiences (iCURE) program. Congrats, Adrianna!
June, 2022. Eugene gave an invited talk and chaired a session at The Post-transcriptional Control of Gene Expression Conference on RNA Decay in Jupiter, Florida. Work on deadenylation kinetics was presented.
June, 2022. Eugene joined the Editorial Board of the Journal of Structural Biology and Journal of Structural Biology: X.
May, 2022. Josh left the lab to begin his graduate studies at Harvard. Thanks, Josh, for your enthusiasm, good humor, and many contributions!
April, 2022. Josh accepted an offer to join the PhD program in the Department of Molecular & Cellular Biology at Harvard University. Congrats, Josh!
April, 2022. Filip Pekovic joined the lab as a postdoc. Filip obtained his PhD from Martin Luther University Halle-Wittenberg in Germany, where he biochemically reconstituted the Drosophila CCR4-NOT complex and studied its regulation by RNA-binding proteins Nanos and Smaug, working with Elmar Wahle. With us, Filip will be working on a collaborative project with Perry Blackshear's group at the NIEHS.
February, 2022. Eugene successfully presented the lab's research accomplishments and plans for the future to the NCI/CCR leadership during the Interim Tenure-Track Review. The concluding assessment from the CCR Director was as follows: "Overall, the review committee was highly impressed with Dr. Valkov’s projects and his progress in establishing an innovative research program. He demonstrated full command of his projects and deep knowledge of the field, which guides many of his projects by identifying the most pressing issues. The questions he is addressing are highly mechanistic, his approaches are cutting-edge and innovative and his experiments are simple, but elegantly designed and impeccably executed. Given that these experiments were done by several members of his lab, he is clearly a strong mentor. Overall, it was felt that Dr. Valkov is well positioned to make meaningful contributions to the RNA processing field and it is clear from his invitations to meetings, and as a reviewer and editor that he is very well respected in the field already. "
December, 2021. Our collaborative study with Georg Stoecklin's group at Heidelberg University, Germany, has been published in Nature Communications. The study describes the functional consequences of the interaction between the RING-type E3 ubiquitin ligase RNF219 and the CCR4-NOT complex. Josh, trained and aided by Yevgen, led the collaboration from our lab and performed all the in vitro assays and protein-protein pulldown assays in the paper.
November, 2021. Wataru was awarded a postdoctoral fellowship from the Japanese Society for the Promotion of Science as part of the Kaitoku-NIH program to support his research. Congrats, Wataru!
November, 2021. Eugene joined the newly established Center for Structural Biology as an Adjunct Investigator. Thank you to Susan Lea for the very kind invitation to collaborate more closely with the Center investigators and participate in its activities, such as the seminar series.
October, 2021. David Turner joined the lab as a postdoc. David obtained his PhD from the University of Cambridge, U.K. He performed his graduate research with Martin Turner at the Babraham Institute, where he designed and generated custom sgRNA libraries targeting RNA-binding proteins to facilitate CRISPR/Cas9-mediated gene knockout screens to identify novel roles for these RBPs. He identified and validated a role for YTHDF2 in promoting terminal B cell differentiation, and in our lab, he will focus on exploring molecular mechanisms of RBPs in mRNA regulation.
August, 2021. Adrianna Hernandez joined the lab as a postbac. She will improve her research skills and then pursue a medical career. The NIDDK provides 50% support for her position, and she will work on a collaborative project with Astrid Haase's group at that institute.
June, 2021. Ilayda Sen, an undergraduate at Ohio State, joined the lab for a month as a virtual summer intern (virtual due to COVID restrictions). She participated in preparing and analyzing RNA sequencing data while working with Khadija.
January, 2021. Leah Pitman joined the lab as a postbac. She will work on a collaborative project with Richard Maraia's group at the NICHD on mRNA regulatory mechanisms and RNA-binding proteins.
June, 2020. Khadija Rebbani joined the lab as a senior postdoc (research fellow). Khadija previously worked at the Cancer Science Institute, part of the National University of Singapore. With us, she will bring her expertise in molecular biology and cellular techniques to help us test mechanistic hypotheses in cells and establish this expertise in the lab. Due to COVID restrictions, Khadija's physical start in the lab has been delayed several months.
April, 2020. Eugene returns to the lab (alone) to produce and purify recombinant SARS CoV-2 Spike protein to support COVID-19 research efforts at the NIH.
March, 2020. Maryland declares a state of emergency and catastrophic health emergency to control and prevent the spread of COVID-19 within the state. The NIH shifts non-mission-critical laboratory operations to a minimal maintenance phase, and a long period of pandemic restrictions and teleworking begins.
February, 2020. Eugene gave an invited seminar at the NIEHS in North Carolina. Thanks to Traci Hall for the very kind invitation and for hosting! I learned much about the wonderful work on that NIH campus and met many new friends and colleagues.
November, 2019. Yevgen Levdansky finally makes his way stateside from Max Planck Institute in Tübingen, Germany, to continue his PhD and begin a new phase of his life and career!
December, 2019. Eric Simko joined the lab as a postdoc. Eric was previously a PhD student at Johns Hopkins University, where he worked with Jiou Wang on the characterization of the interaction between paraspeckle protein NonO and long non-coding RNA Neat1. Eric is interested in the molecular mechanisms regulating the 5' end of the mRNA in our lab.
October, 2019. Josh Corbo joined the lab as a postbac and helped set it up and running!
August, 2019. Eugene arrived in the US and joined the NCI as a Stadtman tenure-track investigator. The Messenger RNA Regulation and Decay section of the RNA Biology Laboratory, a.k.a. the Valkov Lab, is officially inaugurated!
Alumni
Lab Life

The Valkov Lab class of 2024 at Greenbrier State Park.

The Protein Technology Core joined us for our mini-retreat!

Filip, Claire, and Annie.

David, Eric, and Tanner.
Lab retreat at Greenbrier State Park, Maryland, in the spring of 2022.

Lab outing to Brewer's Alley in downtown Frederick to celebrate Yevgen's thesis defense.


Yevgen was very pleased with his Ph.D. hat after the successful defense of his doctoral thesis.





Eugene is part of a sizeable delegation of the NIH intramural community at the annual RNA Society conference in Boulder, Colorado, in June 2022.

Eugene at the FASEB conference on the Post-transcriptional Control of Gene Expression: Mechanisms of RNA Decay in Jupiter, Florida (June 2022).

Late-night discussions at the FASEB meeting in Florida

A 'condensate' of RNA biologists swimming during one of the break sessions at FASEB

Yevgen outside the Moderna headquarters in Boston during a collaborative visit.

Cherry blossom outside our building on Campus in early March 2023.

The fall on campus produces some stunning colors among the trees.
Eugene is giving a talk at UC Riverside in March 2023.