Stefan Ambs, Ph.D., M.P.H.
- Center for Cancer Research
- National Cancer Institute
- Building 37, Room 3050B
- Bethesda, MD 20892-4258
- 240-760-6836
- ambss@mail.nih.gov
RESEARCH SUMMARY
Dr. Ambs conducted integrative epidemiology studies of prostate and breast cancer with a focus on cancer health disparities and risk factors that alter tumor biology. His group utilized epidemiological and translational research and data science approaches to identify exposures and related pathways that promote tumor development and progression and impact patient survival. The team sought to improve disease outcomes among African Americans by gaining insight into their risk factors and tumor biology, and how this knowledge can be harnessed to improve cancer prevention and inform new strategies for clinical management and therapy development. Previous research approaches included the assessment of the tumor transcriptome, epigenome, metabolome, proteome, and mutational signatures, using advanced technologies, and sought to integrate the social environment.
Areas of Expertise
Research
The Molecular Epidemiology Section (MES) brought together a unique skill set that combined epidemiological and laboratory research. We performed molecular epidemiology studies of prostate and breast cancer with a focus on health disparity and utilized translational research and systems biology approaches to identify exposures and related pathways that drove tumor development and progression. We sought to improve disease outcomes among African-Americans by gaining an understanding of their tumor biology, and how this knowledge could be harnessed to improve cancer prevention and care. Our research included the large-scale analysis of gene expression, the evaluation of the tumor metabolome and proteome, and the discovery of biomarkers for disease development and progression. It also involved the analysis of genetic variations and their association with cancer using candidate gene approaches and the application of genome-wide association studies. Thus, our program applied innovative research strategies and was complementary and fully integrated into the transdisciplinary and collaborative research environment within the Laboratory of Human Carcinogenesis (LHC). This environment used shared resources with a common Precision Medicine approach that integrated biology, population-based research, and clinical studies to advance prevention, early diagnosis, and clinical care of common and lethal cancer types.
MES was supported by a resource contract that provided an infrastructure to conduct epidemiological studies in the greater Baltimore area and to collect biospecimens, including tumor and non-tumor tissues and blood samples. The contract was successfully used to collect fresh-frozen breast tumors from African-American (AA) and European-American (EA) patients with long-term survival follow-up. This valuable resource enabled us to recruit AA and EA males into a large case-control study of prostate cancer (976 cases, 1034 population-based controls) to address additional needs for health disparity research. Recruitment into this study was completed in 2016, a major milestone for MES, establishing a cohort with a large biospecimen repository in which half of the cases and controls were self-identified and ancestry-typed AA men. Additionally, we were recruiting breast cancer patients into a research study that would evaluate the relationship between stress-related exposures (e.g., social isolation and discrimination) and tumor biology in breast cancer and compare AA with EA patients in this study.
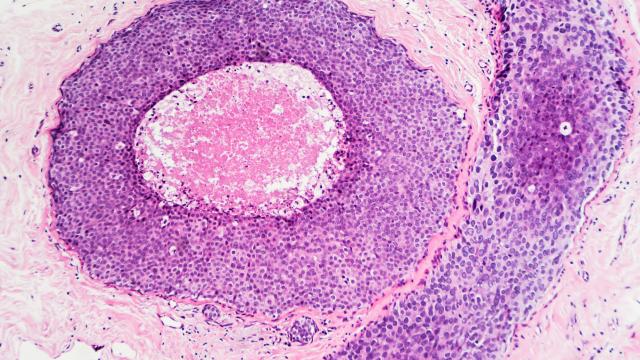
MES strove to use novel and high-risk approaches to gain an understanding of tumor biology in the context of environmental exposures, health disparity, ancestry, and low-penetrance genetic susceptibility. We sought to identify exposures and related pathways that drove tumor development and progression, and then examined how this knowledge could be used to improve cancer prevention and care. One example was the discovery of an immune-inflammation signature in prostate tumors of AA men using transcriptomics and the translation of this finding into the design of a population-based case-control study, which showed that the anti-inflammatory drug, aspirin, might significantly reduce aggressive disease in this patient group. Another example of an innovative design was our breast cancer metabolome study that combined metabolome profiling with a large-scale analysis of gene expression and DNA methylation, leading to the characterization of 2-hydroxyglutarate as a Myc-linked breast cancer oncometabolite and the discovery of a poor outcome tumor subtype with a distinct DNA methylation profile, a high tissue concentration of 2-hydroxyglutarate and heightened occurrence in AA patients.
Our research program was ultimately aimed to improve disease prevention and patient care. The scope of our research program was complementary to other research activities at the LHC and was aligned with the vision of building a collaborative Precision Medicine framework within the LHC and NCI. It was also consistent with the priorities set by the Cancer Moonshot Task Force. We had a strong common interest to improve health outcomes through biomarker discovery and cancer health disparity research. Like other research programs at the LHC, MES continued to examine the contribution of the tumor microenvironment, infections, and chronic inflammation to cancer development and aggressiveness. Our goals were achieved in a collaborative research environment and were consistent with the overall goals of the NCI Intramural Research Program and NCI’s Precision Medicine Initiative for Oncology that sought to advance research in Precision Medicine and Prevention, cancer health disparity and cancer inflammation, focusing on common and lethal cancers. NCI also encouraged an interdisciplinary research approach that led to more effective methods for prevention, early detection, diagnosis, and therapy. These priorities were firmly embedded in our program.
Cancer risk was driven by carcinogen exposure and inherited or acquired host factors. Inherited susceptibility factors (e.g., common variations in the genetic code and inherited mutations within a family), modulated the effects of carcinogen exposure on human cancer risk. Heritable factors could also influence a patient's response to therapy and could become biomarkers for therapeutic outcome. Heritable effects of genes on cancer risk ranged from low to high. Inherited mutations in tumor suppressor genes (e.g., BRCA1, BRCA2, TP53, PTEN) were associated with familial cancer syndromes and predisposed individuals to a high risk of cancer, largely independent of any additional exposure. However, these mutations were rare, and these highly penetrant single gene mutations accounted for less than 10% of all cancers. In contrast, common genetic variations, such as single nucleotide polymorphisms, contributed to common cancers. It was thought that most of cancer cases were caused, at least in part, by the collective effect of multiple, low penetrant genetic variations in a large group of genes and their interactions with environmental exposures. Our group studied common genetic variations in genes of cancer-related pathways (e.g., the inflammation pathway), and examined how these variations influenced cancer risk in the context of ethnicity and/or exposure to environmental risk factors. We were also evaluating how common genetic variations influenced the response of patients to therapy.
The Institute of Medicine concluded that not all segments of the U.S. population had equally benefited from the advances in our knowledge and treatment of cancer. As a result, AAs had the highest death rates from all cancer sites combined, and from malignancies of the lung, colon and rectum, breast, prostate, and the cervix of all racial groups in the United States. Recent research showed that racial disparities in prostate and breast cancer survival between AA and EA patients could persist in randomized clinical trials, indicating that intrinsic differences in tumor biology might have contributed to the disparities in survival between these two patient cohorts. Our group sought to identify the race/ethnic differences in tumor biology that might either influence the presentation of the disease (e.g., causing a more aggressive disease among AA patients), or influence the response to therapy (e.g., causing an inferior response among AA patients). Any of these differences, if they existed, were not likely to be generalizable, meaning that all AA patients were equally different from all EA patients. Rather we believed that certain biologically-derived poor outcome factors were more prevalent among AA patients than EA patients, and that future clinical interventions would have to target these factors across all patient groups.
Acute and chronic inflammation was commonly observed in prostate biopsies and frequently occurred near prostatic intraepithelial neoplasia, a precursor lesion for prostate tumors. It was proposed that the inflammation-to-carcinoma sequence was common in the development of human prostate cancer. A pro-inflammatory tumor environment was also a trigger for disease progression and cancer metastasis. The causes of prostatic inflammation were poorly understood but could have been caused by chronic infections and other environmental exposures. Our research discovered a pro-inflammatory interferon gene signature in tumors of AA patients, suggesting the possible involvement of viral infections in disease pathology. We also observed a unique gene expression profile in prostate tumors associated with current smoking that indicated an altered immune environment in tumors of current smokers. This observation could have been very significant because current smoking increased the risk of metastatic prostate cancer. These findings were further pursued in the context of the NCI-Maryland Case-Control study.
Publications
- Neighborhood Deprivation and DNA Methylation and Expression of Cancer Genes in Breast Tumors. JAMA Netw Open 6: e2341651,2023
- Diabetes-associated breast cancer is molecularly distinct and shows a DNA damage repair deficiency. JCI Insight,8: e170105,2023
- Circulating trans fatty acids are associated with prostate cancer in Ghanaian and American men. Nat Commun., 14: 4322,2023
- Population-specific Mutation Patterns in Breast Tumors from African American, European American, and Kenyan Patients. CRC 2023
- Chromatin accessibility landscape of human TNBC cell lines reveals variation by patient donor ancestry. CRC 2023
- Association of Neighborhood Deprivation With Prostate Cancer and Immune Markers in AA and EA Men. JAMA Netw Open 2023
- Serum proteomics links suppression of tumor immunity to ancestry and lethal prostate cancer. Nat Commun., 13: 1759,2022
MYC-driven accumulation of 2-hydroxyglutarate is associated with breast cancer prognosis
Liver- and Microbiome-derived Bile Acids Accumulate in Human Breast Tumors and Inhibit Growth and Improve Patient Survival.
Urinary Thromboxane B2 and Lethal Prostate Cancer in African American Men.
Integrated proteotranscriptomics of breast cancer reveals globally increased protein-mRNA concordance associated with subtypes and survival.
IFNL4-ΔG Allele Is Associated with an Interferon Signature in Tumors and Survival of African-American Men with Prostate Cancer.
Biography
Stefan Ambs, Ph.D., M.P.H.
Dr. Stefan Ambs earned a Master's degree in Biochemistry from the University of Tuebingen (1988) and a Master of Public Health degree (Epidemiology) from the Johns Hopkins Bloomberg School of Public Health (2005). He completed his Ph.D. thesis (1992) at the Institute of Toxicology, University of Wuerzburg, Germany, and was trained as a Postdoctoral Fellow at the Laboratory of Human Carcinogenesis (1992-1997), National Cancer Institute, Bethesda, Maryland. He continued his research at a biotechnology company in California and at the Aventis Genomics Center in Cambridge, Massachusetts. Dr. Ambs joined the NCI as a tenure-track investigator in November of 2001. He became a tenured Senior Investigator in 2010.
Dr. Stefan Ambs retired from CCR in May of 2025.
Covers
ADHFE1 is a breast cancer oncogene and induces metabolic reprogramming
The cover image depicts the role of mitochondrial ADHFE1 in D-2-hydroxyglutarate production, highlighting the contributions of the enzyme and oncometabolite to breast cancer progression.
Metabolic reprogramming in breast tumors is linked to increases in putative oncogenic metabolites that may contribute to malignant transformation. We previously showed that accumulation of the oncometabolite, 2-hydroxyglutarate (2HG), in breast tumors was associated with MYC signaling, but not with isocitrate dehydrogenase (IDH) mutations, suggesting a distinct mechanism for increased 2HG in breast cancer. Here, we determined that D-2HG is the predominant enantiomer in human breast tumors and show that the D-2HG–producing mitochondrial enzyme, alcohol dehydrogenase, iron-containing protein 1 (ADHFE1), is a breast cancer oncogene that decreases patient survival. We found that MYC upregulates ADHFE1 through changes in iron metabolism while coexpression of both ADHFE1 and MYC strongly enhanced orthotopic tumor growth in MCF7 cells. Moreover, ADHFE1 promoted metabolic reprogramming with increased formation of D-2HG and reactive oxygen, a reductive glutamine metabolism, and modifications of the epigenetic landscape, leading to cellular dedifferentiation, enhanced mesenchymal transition, and phenocopying alterations that occur with high D-2HG levels in cancer cells with IDH mutations. Together, our data support the hypothesis that ADHFE1 and MYC signaling contribute to D-2HG accumulation in breast tumors and show that D-2HG is an oncogenic metabolite and potential driver of disease progression.
ADHFE1 is a breast cancer oncogene and induces metabolic reprogramming. Mishra P, Tang W, Putluri V, Dorsey TH, Jin F, Wang F, Zhu D, Amable L, Deng T, Zhang S, Killian JK, Wang Y, Minas TZ, Yfantis HG, Lee DH, Sreekumar A, Bustin M, Liu W, Putluri N, and Ambs S. J Clin Invest. 128(1):323-340, 2018.