Sandra L. Wolin, M.D., Ph.D.
- Center for Cancer Research
- National Cancer Institute
- Building 560, Room 11-82C
- Frederick, MD 21702-1201
- 301-846-1237
- wolinsl@nih.gov
RESEARCH SUMMARY
Sandra Wolin studies the biogenesis, function and turnover of RNA molecules. Her laboratory has identified proteins that recognize misfolded and otherwise defective RNAs. By studying a bacterial ortholog of one such protein, the ring-shaped Ro60 autoantigen, they discovered that this protein is tethered by noncoding “Y RNA” to a ring-shaped nuclease, forming a double-ringed ribonucleoprotein machine specialized for structured RNA degradation. The laboratory is characterizing this new RNA degradation machine, identifying additional roles for Ro60 and Y RNAs in both human cells and bacteria, and uncovering other pathways by which defective and damaged RNAs are recognized and handled. Her laboratory also investigates how failure to degrade these RNAs contributes to human disease.
Areas of Expertise

Sandra L. Wolin, M.D., Ph.D.
Research
Our laboratory studies how noncoding RNAs function, how cells recognize and degrade defective RNAs, and how failure to degrade these RNAs affects cell function and contributes to human disease. Most cellular RNA does not encode proteins, and truncated, misfolded and aberrant RNAs can accumulate due to mutations, transcriptional errors and processing mistakes. Some forms of environmental stress, such as exposure to oxidants and ultraviolet light, result in RNA damage, yet little is known of how damaged RNAs are recognized and handled.
One pathway that we study involves noncoding RNA-protein complexes known as Ro60 ribonucleoproteins (RNPs). These RNPs are clinically important targets of the immune system in patients suffering from two rheumatic diseases, systemic lupus erythematosus and Sjögren’s syndrome. The major protein component, the ring-shaped Ro60 autoantigen, is present in most animal cells and ~5% of bacteria. In all studied organisms, Ro60 binds noncoding RNAs called Y RNAs. By studying Ro60 in bacteria, we discovered that this protein is tethered by Y RNA to a ring-shaped nuclease, forming a double-ringed RNP machine specialized for structured RNA degradation. We showed that Ro60 contributes to survival of both mammalian cells and bacteria in the presence of stresses, such as ultraviolet light, that damage nucleic acids. We also provided evidence that bacterial Ro60 RNPs may act through molecular mimicry to trigger autoantibody formation in susceptible patients. Our current goals are to uncover additional roles for Ro60 and Y RNAs in both mammalian cells and bacteria. We are also studying ways in which this and other RNA surveillance pathways contribute to normal cell physiology and prevent diseases such as cancer.
Relevance to cancer
While much is known as to how DNA damage contributes to carcinogenesis, less is known about the effects of RNA damage. Many environmental agents that damage DNA also affect RNA integrity and alter RNA pools. For example, UV irradiation causes RNA-protein crosslinks and increases transcription of retrotransposons. Many common chemotherapeutics, such as mitomycin C and bleomycin, also damage RNA. By uncovering novel mechanisms by which cells deal with the effects of RNA damage, our work could advance understanding of the paths that lead to cancer and increase knowledge as to how chemotherapeutic agents function.
Publications
Noncoding Y RNAs Regulate the Levels, Subcellular Distribution and Protein Interactions of their Ro60 Autoantigen Partner
Cellular RNA Surveillance in Health and Disease
Biography

Sandra L. Wolin, M.D., Ph.D.
Dr. Wolin earned her A.B. in Biochemical Sciences from Princeton University, her M.D. from the Yale School of Medicine and her Ph.D. from the Department of Molecular Biophysics and Biochemistry at Yale University. After postdoctoral training at the University of California, San Francisco, she returned to the Yale School of Medicine as an Assistant Professor and rose to the rank of Professor in the Departments of Cell Biology and Molecular Biophysics and Biochemistry. From 2014-2017, she served as Director of the Yale Center for RNA Science and Medicine. She joined the NCI in 2017 as the inaugural Chief of the newly formed RNA Biology Laboratory and Head of the NCI RNA Biology Initiative. She is an elected Fellow of the American Association for the Advancement of Science (2013), the American Academy of Microbiology (2014), the American Academy of Arts and Sciences (2023) and the National Academy of Sciences (2024).
Job Vacancies
| Position | Degree Required | Contact Name | Contact Email |
|---|---|---|---|
| Postdoctoral Fellow - RNA biology, noncoding RNAs and RNPs | Ph.D. or equivalent | Sandra Wolin | sandra.wolin@nih.gov |
Team
News
April 2024: Congratulations to Sandy on her election to the National Academy of Sciences!
July 2023: Congratulations to Hyeyeon for winning a 2024 Fellows Award for Research Excellence (FARE)!
April 2023: Sandy is elected to the American Academy of Arts and Sciences.
October 2022: Congratulations to Sandra Williams for receiving a best poster award at the NCI RNA Biology Initiative Retreat!
June 2022: Congratulations to Sandy on her election as RNA Society President!
December 2021: Sandy receives the ASCB Sandra Mazur Senior Leadership Award
April 2021: Congratulations to Xinguo for winning first place and a travel award for his oral presentation at the 2021 Staff Scientists/Staff Clinicians retreat!
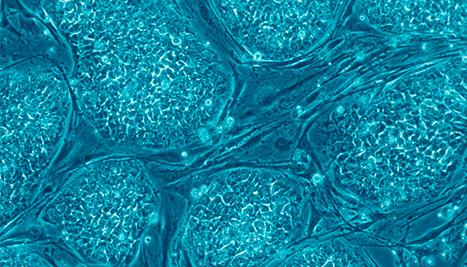
New mechanism for regulating differentiation discovered in human embryonic stem cells
Human embryonic stem cells.
Photo credit: Wikimedia Commons
CCR researchers have found that the RNA exosome plays an important role in restraining differentiation of human embryonic stem cells (ESCs). Differentiation is the process by which human ESCs cells develop into the three germ layers that form the body: endoderm, mesoderm and ectoderm. All tissues and organs in the body can be formed from human ESCs.
This laboratory research is the first time that the RNA exosome has been studied in human ESCs as opposed to progenitor cells, which are more mature cells that have already begun to differentiate into a specific cell type. This study, led by Sandra L. Wolin, M.D., Ph.D., Chief of the RNA Biology Laboratory, appeared July 15, 2019, in the Journal of Cell Biology.
June 2019: Congratulations to Yuanyuan for winning a 2020 FARE Award.
Kevin is selected for the NCI iCURE program!
New study shows normally helpful natural bacteria may also trigger lupus
CCR scientists have discovered that a protein produced by bacteria that naturally inhabit our bodies may trigger the autoimmune disease lupus. The results of the study could unveil an entirely new set of drug targets for treating lupus and other autoimmune diseases. Read more…
July 2017: Cedric has won a FARE Award. Congratulations, Cedric!
Covers

Commensal orthologs of the human autoantigen Ro60 as triggers of autoimmunity in lupus
Lupus: No Longer a Lone Wolf. Lupus (represented by the wolf, Canis lupus) is a chronic autoimmune disorder that progresses over decades. Greiling et al. analyzed commensal bacteria in lupus patients and identified species with Ro60 proteins similar to human Ro60, an early autoantigen in lupus. Bacterial Ro60 could activate patient lymphocytes, and colonization of mice with the bacterium instigated lupus-like symptoms. These results suggest that immune recognition of a bacterial protein leads to cross-reactive immune cells targeting the human version, thereby initiating lupus.
Greiling et al., Science Translational Medicine, Vol 10, Issue 434, 28 March 2018: Vol 10, Issue 434
Lab Life

Celebrating at the RBL Holiday Party

Celebrating RNA Day at the NCI RNA Biology Initiative picnic

Farewell lunch for Marco

Celebrating Sandy's election to the American Academy of Sciences

Hiking in Greenbrier State Park