HeLa cell
Image courtesy of NIH Flickr
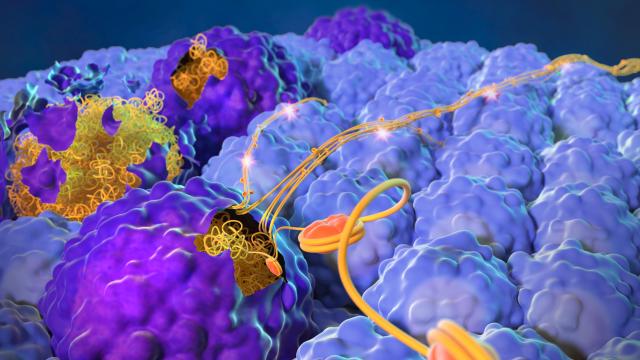
Acyl-coenzyme As (CoAs) are a group of small molecule metabolites that are the building blocks and chemical energy source for all living organisms. Recent evidence indicates that metabolites such as acyl-CoAs can also interact with proteins to drive the development of cancer. The transient nature of acyl-CoA binding, however, has prevented detection of these events by traditional methods.
CCR researchers have developed an assay known as CATNIP (CoA/AcetylTraNsferase Interaction Profiling) to get a better handle on how acyl-CoAs directly interact with proteins and fuel malignancy. Jordan L. Meier, Ph.D., Senior Investigator in the Chemical Biology Laboratory, working in collaboration with Michael Washburn, Ph.D., Stowers Institute for Medical Research, Kansas City, MO, have used this assay to construct one of the first comprehensive maps of cellular metabolite/protein interactions.
The CATNIP method is unique in that it uses an approach known as chemoproteomics to look upstream at the source of metabolic signaling, identifying proteins that are physically bound by metabolites. This is in contrast to earlier approaches which used downstream techniques, such as gene expression profiling, to determine signaling mechanisms altered by metabolites in cancer.
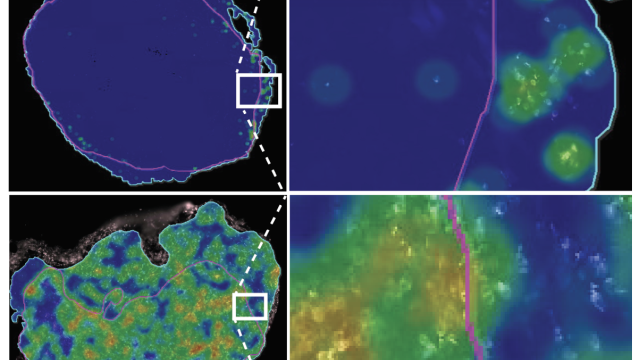
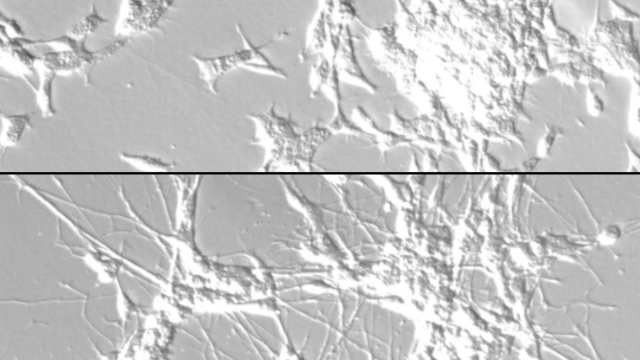
The researcher’s study, published December 10, 2019, in Cell Chemical Biology, used well-known ovarian cancer cells (HeLa cells), which grow quickly, allowing the scientists to obtain large amounts of cancer-derived protein. These proteins are found not only in ovarian cancer but also in many other types of cancer, so the investigators believe that they will be able to better understand fundamental principles of how acyl-CoA and proteins interact.
The insights gained by studying acyl-CoAs are vital because they play a unique role in almost every biochemical pathway in the cell; because they are ancient and are used by all types of organisms; because many different types of enzymes interact with them, giving them the potential to affect many different types of signaling pathways; and because they directly feed histone acetylation, a signaling pathway that increases gene expression and has been implicated in cancer.
With the use of CATNIP, the investigators have learned that acyl-CoAs potentially regulate enzymes on a larger scale than was previously thought, noted Meier. He also said that his team was surprised that most enzymes bind to multiple acyl-CoAs, not just to their known substrates.
Dr. Meier believes that the CATNIP assay could potentially map acyl-CoA/protein interactions in living organisms, and this is one of the future directions they envision for this technology. They also hope to better understand how acyl-CoA/protein interactions are remodeled during transformation, e.g., cancer development, which could lead to the identification of signaling hubs to target for therapeutic intervention.
Ultimately, the scientists believe that the CATNIP method could have valuable applications in drug discovery by helping to understand the selectivity and affinity of newly developed inhibitors of histone acetylation as well as to spur researchers to think about new ways to target these processes by altering cellular metabolism.