miRNAs, lncRNAs and circRNAs are newfound types of non-coding RNAs that are shedding light on the regulation of gene expression.
Photo courtesy of the NIH IRP
MicroRNAs have an enormous influence over what happens inside cells. By blocking the activity of specific sets of genes, they help control virtually every known biological pathway and process. Disruptions in microRNAs have been linked to many diseases, and understanding how these molecules function, which genes they control and how they themselves are regulated are high priorities in cancer research.
New research from Shuo Gu, Ph.D., Investigator in the RNA Biology Laboratory, shows that when a microRNA undergoes a common modification called uridylation, its genetic targets change. A single microRNA can regulate hundreds of different genes, but when a microRNA is uridylated, the team found, even more genes come under its control.
Modifications to microRNAs that either clip off or add to their short sequences are widespread, but it has not been know what effect these modifications have on microRNA function. Until the new study, reported June 6, 2019, in Molecular Cell, there were few clues to suggest that uridylation might alter the way a microRNA interacts with its targets.
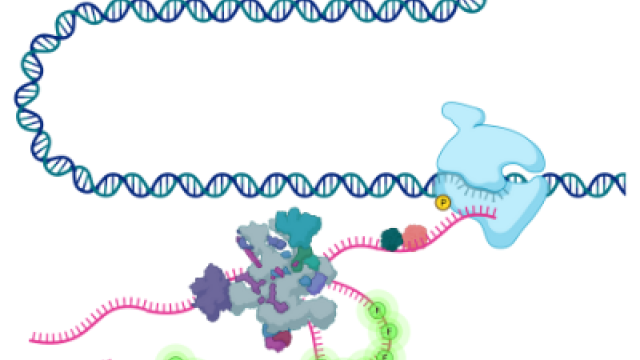
MicroRNAs block the activity of specific genes by binding to the messenger RNAs that relay a gene’s information to the cell’s protein-making machinery. To recognize its targets, each microRNA relies on a stretch of seven nucleotides (genetic letters) within its own sequence that binds to a complementary sequence in messenger RNA, a segment known as the microRNA’s seed.
Uridylation, which adds a string of Us to the 22-nucleotide sequence that makes up a microRNA, does not alter the seed, so researchers had not expected it to impact the gene regulators’ target recognition. Instead, the modification was thought to be primarily a way for cells to control the stability and abundance of specific RNAs. Still, Gu says, changes in the levels of sequence-modified microRNAs have been linked to cancer, suggesting they may play an important role in the disease process.
Gu’s team tested several uridylated microRNAs and found that each was able to repress mRNA targets that the unmodified microRNAs could not. Their experiments with a microRNA known as miR-27a revealed that the modified microRNA’s tail facilitated pairing with targets that were poor matches for the seed.
Based on the features they determined were necessary for this tail-assisted pairing, the team predicted that uridylation could allow a microRNA to interact with hundreds of new genetic targets. Now, they plan to explore how healthy cells use this shift to control normal processes as well as uridylated microRNAs’ role in disease.