Stavroula Mili, Ph.D.
- Center for Cancer Research
- National Cancer Institute
- Building 37, Room 2042
- Bethesda, MD 20892-4256
- 240-760-6844
- voula.mili@nih.gov
RESEARCH SUMMARY
Our research program focuses on the roles of localized RNAs in mammalian physiology. Our work spearheaded the study of RNAs targeted to protrusive regions of cells and established that protrusion localized mRNAs are functionally important during cell migration and cancer cell invasion. This mRNA localization mechanism is further linked to physiologically and clinically relevant processes since it occurs in vivo and is deregulated by pathogenic mutations. We use a variety of approaches including single-molecule RNA imaging in fixed and live cells, bioengineering methods as well as 3D and in vivo models. Our goal is to understand the molecular regulation, functional consequences and disease associations of localized RNAs.
Areas of Expertise

Stavroula Mili, Ph.D.
Research
Regulation and Functions of Localized RNAs in Non-Neuronal Cells
Active targeting of mRNAs to specific regions in the cytoplasm is a prevalent and physiologically important phenomenon. Localized RNAs have been studied predominantly in highly polarized cells of model organisms and in mammalian neurons, where they have essential roles in development, cell fate determination and neuronal function. By contrast, localized RNAs and the functional roles they exert in less polarized and more dynamic somatic cells have been until recently largely unexplored. Our research program is leading efforts in this understudied area, focusing on mRNAs localized to protrusive regions of cells. We have shown that these RNA localization events are important for mammalian cell migration and cancer invasion and have linked them to physiologically and clinically relevant processes. We are trying to comprehensively delineate on a molecular level the regulation and functional consequences of RNA localization and localized translation. We are further incorporating our findings into increasingly physiologically relevant systems of study, such as multicellular 3D cultures, mechanically diverse contexts, and in vivo models.
Available positions
Our group has openings for motivated post-doctoral and post-bac fellows who wish to study the molecular regulation and functional roles of localized RNAs. If you are interested in our work, please send your C.V., a statement of research interests and the names and contact details of 3 references to voula.mili@nih.gov
Specific research directions that we are pursuing include:
Cis-elements, trans-acting factors and mechanisms controlling RNA localization at protrusions
Using a combination of single-molecule live-cell RNA imaging, biochemical and genetic approaches, we have been dissecting the molecular players involved in localizing RNAs to cell protrusions and in regulating their translation. These factors include cis-acting elements (GA-rich regions) present in the 3’ UTR of transported RNAs; a specific subset of modified, detyrosinated microtubules; the microtubule-based motor KIF1C; and various RNA-binding proteins. Most notably we identified the tumor suppressor protein APC and the neurodegeneration related protein FUS as novel regulators of a specific subset of protrusion localized RNAs, which we termed ‘APC-dependent’. (Mili et al, 2008, Nature; Yasuda et al., 2013, J Cell Biol; Moissoglu et al., 2020, EMBO J; Pichon et al., 2020, BioRxiv).
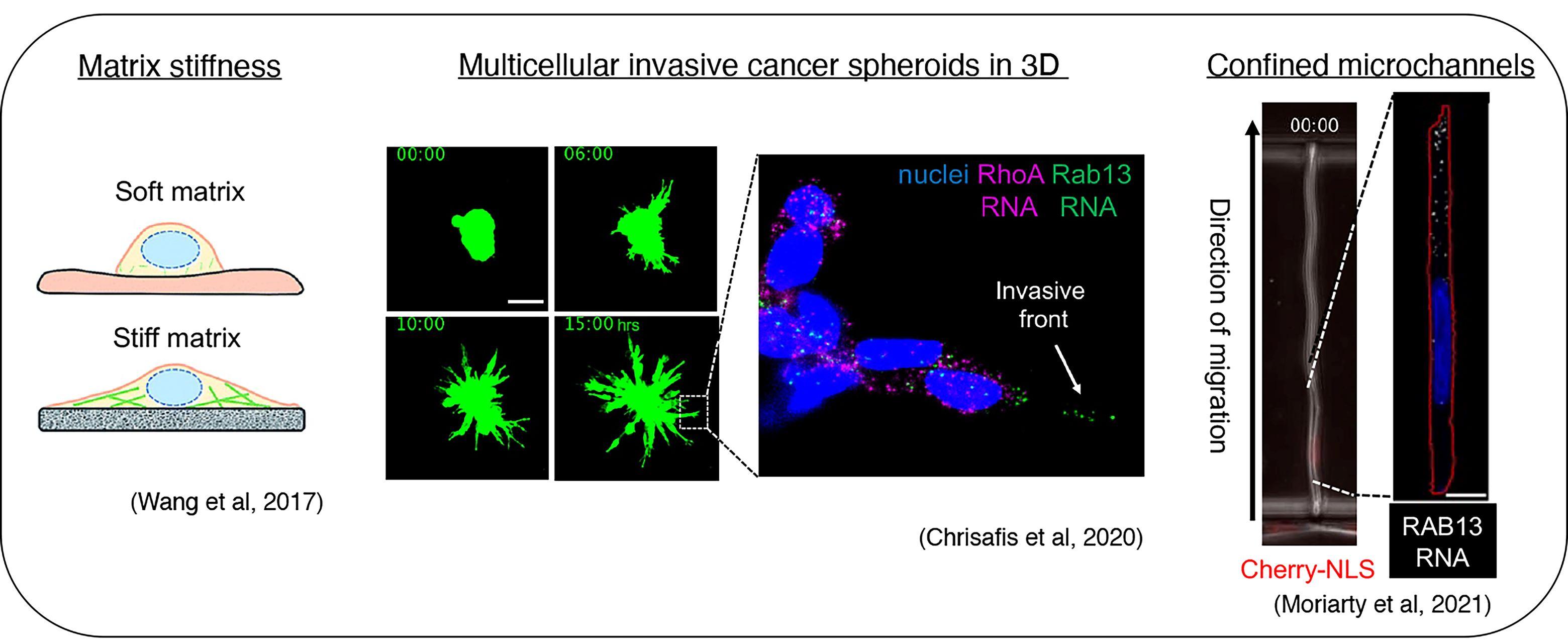
Importantly, we revealed that localization of RNAs at protrusions is under mechanical control and responds to the properties of the extracellular environment. We have been exploring this regulation in simpler 2D culture systems, in multicellular 3D invasive cancer spheroids, and during migration in confined spaces (Wang et al., 2017, Nat. Commun; Chrisafis et al., 2020, PNAS; Moriarty et al., 2021). These findings suggest that, in physiological settings, the specific mechanical properties of the cells and of the microenvironment could impact the regulation and functional outcomes of localized RNAs.
Translational regulation of protrusion-localized RNAs
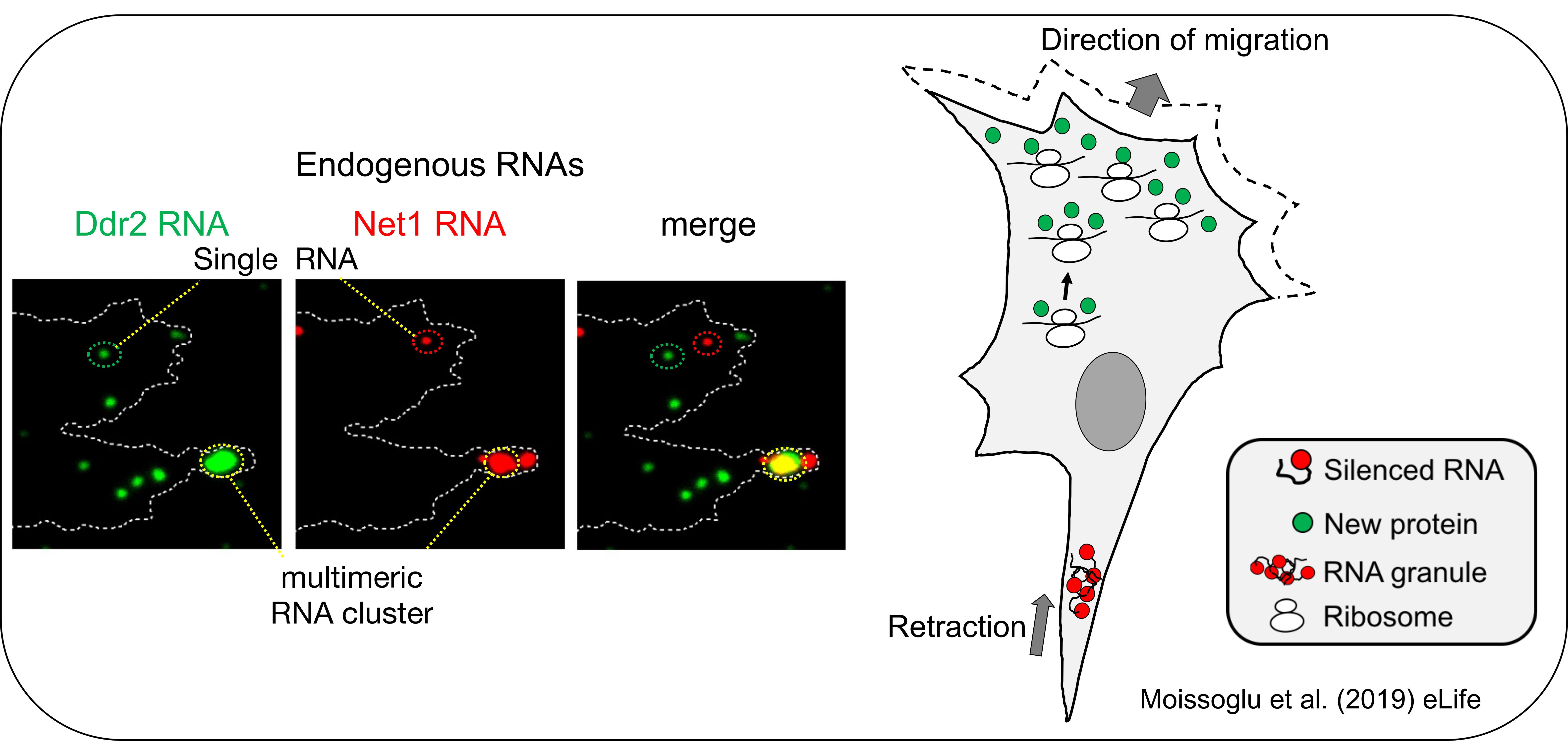
We have shown that protrusion localized transcripts are regulated in novel ways. They exist in two distinct physical states, either as single RNAs or as multimeric clusters composed of multiple RNA species. These multimeric clusters are transiently formed at retracting protrusions of migrating cells. Interestingly, cluster formation is coupled to changes in the translation of the RNAs, with RNAs within clusters being translationally silent (Moissoglu et al., 2019, eLife). These clusters are reminiscent of RNA granules formed by liquid-liquid phase separation. These findings thus reveal a novel mode of dynamic control of translation in migrating cells. They further suggest that the assembly/disassembly of translationally silent RNA clusters can be locally controlled and coordinated with specific cellular behaviors. We continue exploring the factors and signaling events underlying this intriguing local regulation.
Functional consequences of local RNA translation
An important unanswered question in the field concerns how, on a molecular level, localized RNAs exert their functional roles. In large, stably polarized cells, such as neurons or oocytes, the role of RNA localization is thought to be to spatially restrict protein synthesis leading to persistent or transient changes in local protein concentrations. This model is however insufficient to explain the role of localized RNAs in dynamic migrating cells due to their small size, fast rate of diffusion and rapid changes in overall morphology and polarization. Indeed, in non-neuronal cells RNA localization often fails to correlate with protein enrichment, suggesting that for many transcripts there likely exist additional unappreciated ways by which RNA localization controls gene expression.
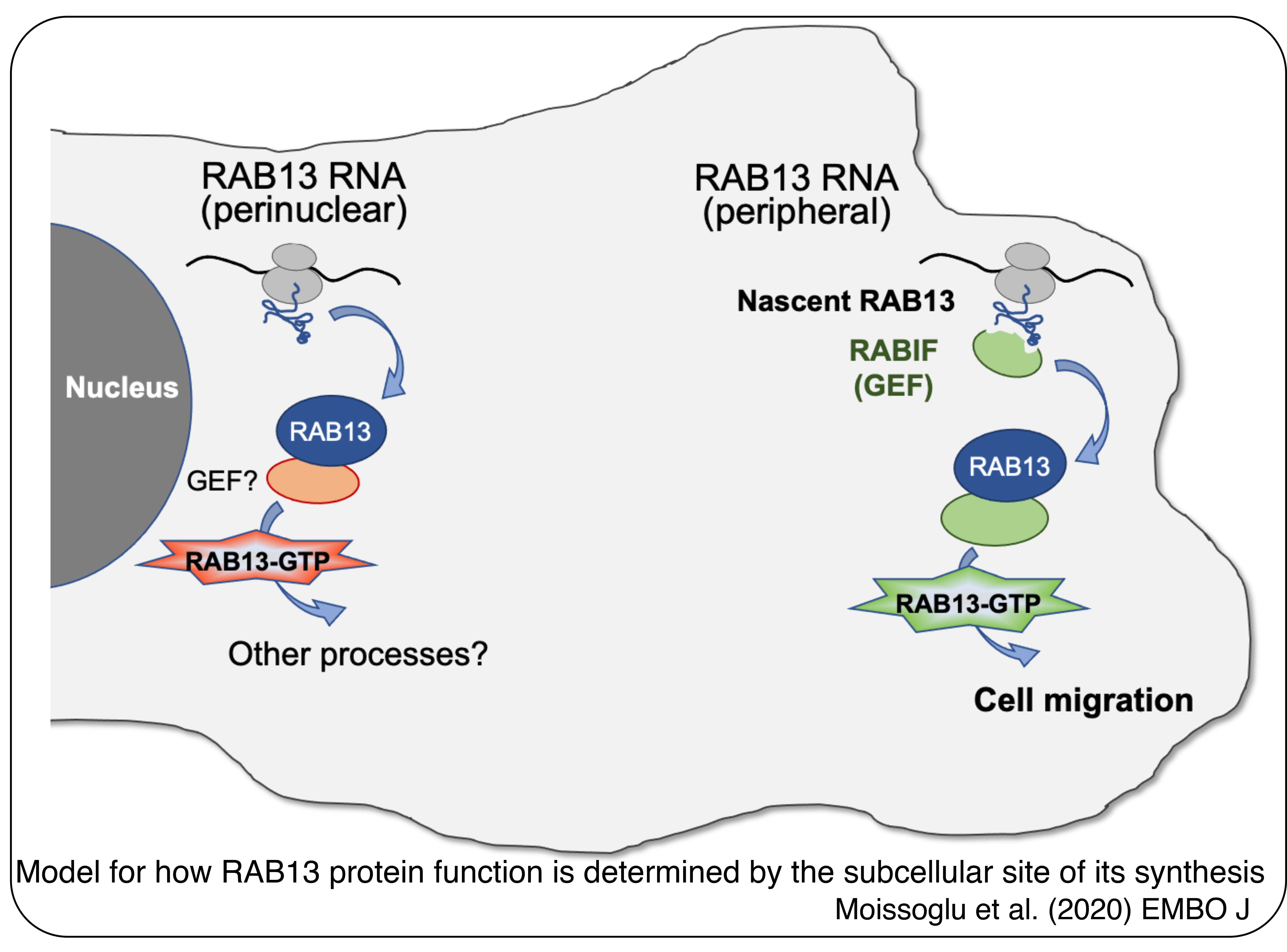
Our studies have provided important insights into this question. We have used as a case study the RAB13 RNA, which encodes a GTPase involved in vesicle trafficking. We find that the particular subcellular site of RAB13 protein synthesis can critically affect the ability of RAB13 to support cell migration. On a molecular level, a co-translational interaction of nascent RAB13 with its activator RABIF at the periphery is required for directing the RAB13 GTPase activity to promote cell migration. This occurs without a detectable effect on RAB13 protein distribution (Moissoglu et al., 2020, EMBO J). These findings have provided a molecular explanation for how RNA localization can be important in dynamic cells even without directing local protein accumulation. Our model suggests that specific co-translational interactions in different local environments direct the newly synthesized proteins towards distinct functional outputs. We are extending these findings using different case studies and are examining their broader implications regarding the regulation of protein function.
Contributions of protrusion-localized RNAs during in vivo cancer invasion
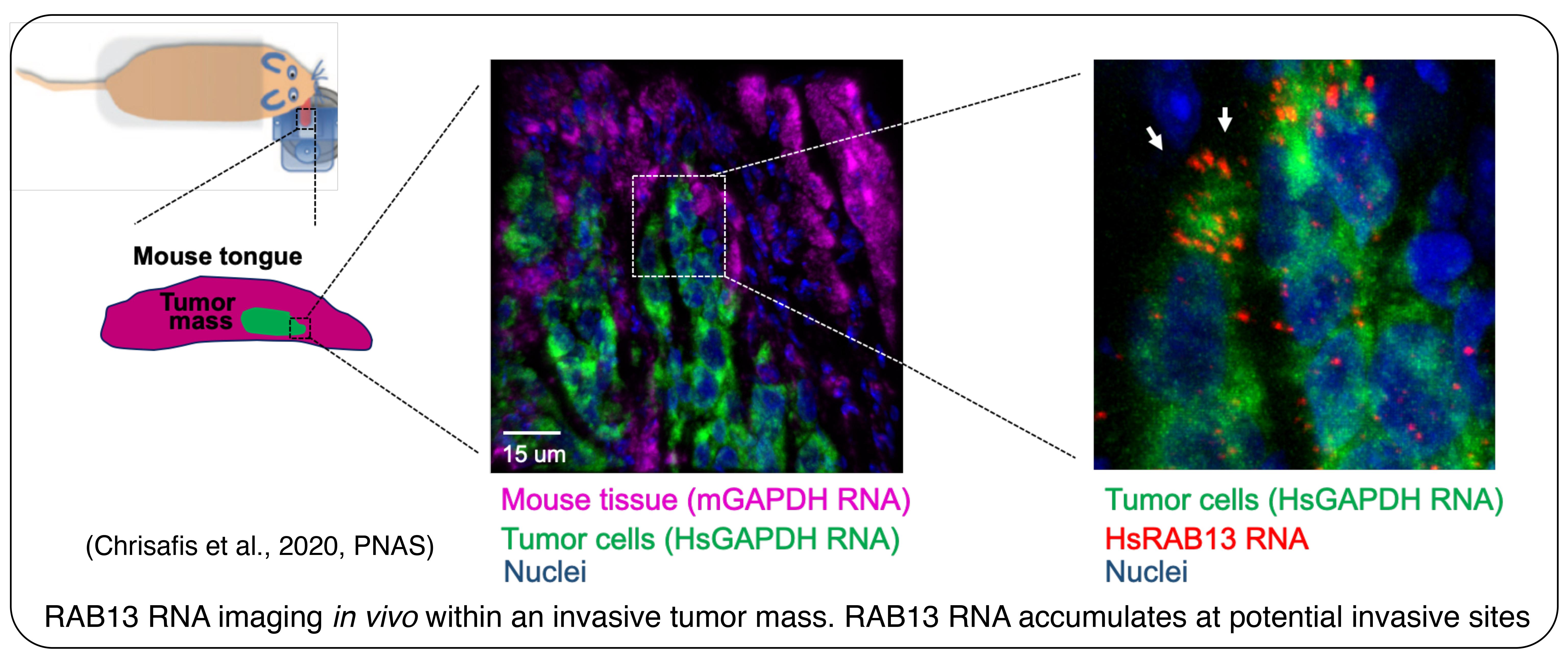
We have exploited our knowledge of the underlying mechanisms to develop antisense oligonucleotides that interfere specifically with localization of particular endogenous transcripts. Using this methodology, which can be applied both in vitro and in vivo, we have demonstrated a requirement for specific localized RNAs not only during migration of individual cells, but also during collective invasion of 3-dimensional multicellular cancer spheroids (Moissoglu et al., 2020, EMBO J; Chrisafis et al., 2020, PNAS). We are extending our findings using xenograft mouse tumor models of cancer invasion.
Deregulation of localized RNAs by pathologic RNA inclusions
The RNA-binding protein FUS associates with RNAs at protrusions and likely regulates their translation (Yasuda et al., 2013, J Cell Biol). FUS belongs to a group of proteins with low-complexity, prion-like domains, which drive formation of membrane-less inclusions/granules, through a process of liquid-liquid phase separation. Inclusion formation by FUS is enhanced by the presence of mutations associated with ALS (amyotrophic lateral sclerosis). We found that cytoplasmic inclusions formed by ALS-FUS mutants preferentially recruit APC-associated RNAs and disrupts their localization at protrusions. Interestingly, RNAs are not silenced within FUS inclusions, but are being actively translated (Yasuda et al., 2013, J Cell Biol). Formation of pathogenic FUS inclusions disrupts localization of APC-dependent RNAs by sequestering the kinesin-1 motor protein and inhibiting formation of detyrosinated microtubules (Yasuda et al., 2017, J Cell Biol). These results highlight a potential role for this pathway in maintaining normal neuronal function. We are interested in understanding the interplay between RNA granule formation and RNA translation and the functional consequences of physiologic or pathologic RNA granules.
Publications
- Bibliography Link
- View Dr. Mili's PubMed Summary.
mRNA location and translation rate determine protein targeting to dual destinations
Collective cancer cell invasion requires RNA accumulation at the invasive front
RNA localization and co-translational interactions control RAB13 GTPase function and cell migration
Translational regulation of protrusion-localized RNAs involves silencing and clustering after transport
Extracellular matrix stiffness and cell contractility control RNA localization to promote cell migration
Biography

Stavroula Mili, Ph.D.
Stavroula (Voula) Mili obtained her BSc degree in Biology from the University of Athens, Greece and her PhD degree in Biomedical Sciences from the Mount Sinai School of Medicine of New York University under Dr. Serafin Pinol-Roma. For her post-doctoral training she joined Dr. Joan Steitz's lab at Yale University and subsequently Dr. Ian Macara's lab at the University of Virginia. During that time, she discovered a localization pathway that targets mRNAs to cellular protrusions. This line of research formed the basis of her independent research group at the NIH. She joined the Laboratory of Cellular and Molecular Biology at the NCI in September 2012 as a Stadtman Investigator and was awarded tenure at NIH in 2022. Research in her lab employs multi-disciplinary approaches to understand the regulation and biological roles of protrusion-localized mRNAs, with an emphasis on using advanced microscopy and diverse in vitro and in vivo mammalian model systems to explore the contributions of localized mRNAs in normal physiology and cancer invasion.
Job Vacancies
We have no open positions in our group at this time, please check back later.
To see all available positions at CCR, take a look at our Careers page. You can also subscribe to receive CCR's latest job and training opportunities in your inbox.