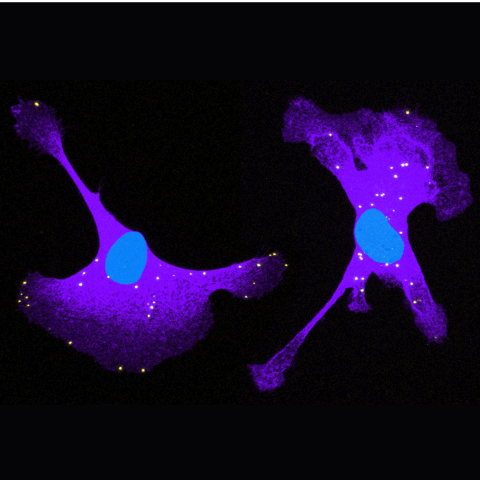
In this image, NET1 mRNA molecules (yellow dots) are shown in breast cancer cells (cell body in purple and nucleus in blue). The researchers used antisense oligos to redistribute the mRNA molecules in the cell to the right. By altering the location of the NET1 mRNA, and thus the site of NET1 protein synthesis, the team could study how location affects the properties of the encoded NET1 protein and the ability of cancer cells to migrate. Image credit: Alexander Gasparski, Ph.D.
Researchers at CCR have discovered two key underlying factors for how the same protein can choose between binding partners and have very different functions within a cell: the location and rate of messenger RNA (mRNA) translation can both determine the fate of a protein. The results, described in a study published in Molecular Cell on July 27, 2023, could lead to more precise control over protein function in cells for therapeutic purposes.
Our cells contain strands of mRNA that encode instructions for producing specific proteins. But researchers have observed that the same protein can exert different functions by binding to different partners. How these partners are selected is not entirely known.
Stavroula Mili, Ph.D., a Senior Investigator in the Laboratory of Cellular and Molecular Biology, and her colleagues have been studying a protein called NET1, which can be found in two areas of the cell. The researchers noticed that the mRNA that codes for NET1 can be found either near the nucleus, at the center of the cell, or near the cell’s surface. They devised ways to alter the distribution of the mRNA to understand its role.
In a series of experiments in cultured cancer cells, they showed that when the mRNA for NET1 was closer to the nucleus, the NET1 protein interacted more with a protein responsible for carrying it into the nucleus. However, when the mRNA was located further away from the nucleus, the NET1 protein interacted more with a different partner that kept it outside of the nucleus, where it helped to enhance the ability of the cancer cells to migrate.
The researchers also noticed that these partners interacted with different regions of the NET1 protein. To shed light on this observation, they manipulated how quickly these regions appeared as the NET1 protein was produced, by changing the rate at which the mRNA is translated into the protein. Strikingly, slow translation favored the partner that interacts with an early-appearing region, while faster translation favored the other partner that interacts with a later-appearing region of NET1. The team’s finding confirms that the rate of mRNA translation also influences the selection of partners and the function of the protein.
Essentially, the microenvironment where protein synthesis takes place and how quickly different parts of a protein emerge during its production significantly influence and determine protein function.
“What was surprising to me was how effective these mechanisms are in terms of influencing one function of the protein or another,” says Mili.
Notably, understanding these differences in protein function could help scientists more precisely manipulate the inner workings of cells for therapeutic purposes. “For example, if a protein has two functions, you could prevent one function, while preserving the other, which could be useful for therapeutic purposes,” explains Mili.
She says that her team suspects similar factors of the cellular microenvironment and translation rates may influence the function of other proteins as well, which they plan to explore in future research.