Colin C. C. Wu, Ph.D.
- Center for Cancer Research
- National Cancer Institute
- Building 560, Room 21-102C
- Frederick, MD, 21702-1201
- 301.846.7683
- colin.wu2@nih.gov
RESEARCH SUMMARY
We are exploring the role of the ribosome in stress response signaling pathways and translational regulation in mammalian systems. To this end, we have developed an improved ribosome footprint profiling methodology that allows us to monitor both the positions and the in vivo functional states of individual ribosomes. Most projects in the laboratory incorporate high-throughput genome-wide approaches, computational tools, and biochemical approaches to define the molecular mechanisms of these pathways in vivo. Our long-term goal aims to congregate actionable insights that can be used to guide the prevention and treatment of human diseases.
Areas of Expertise

Colin C. C. Wu, Ph.D.
Research
We are actively recruiting highly motivated postdoctoral researchers to join our team. If you’re passionate about RNA biology, ribosome quality control, or related areas, please email Colin Wu (colin.wu2@nih.gov).
Our laboratory is broadly interested in translational regulation during cellular stress, especially by the ribosome, ribosome-associated factors, and RNA binding proteins. We employ an integrated approach, combing mass spectrometry, CRISPR screens, high-throughput chemical probing, ribosome profiling, biochemical techniques, and computational tools to identify and characterize novel factors involved in stress sensing and fine-tuning the translational output.
Protein synthesis is essential in all cells. As such, a continuous flow of ribosomes needs to be maintained to support a healthy proteome. Ribosome stalling occurs under a variety of cellular stress conditions, including nutrient deprivation, hypoxia, and oxidative stress. We recently discovered that ribosome stalling triggers 40S subunit decay through a highly choreographed ribosome quality control pathway (Coria, Shah et al., Mol Cell 2025).
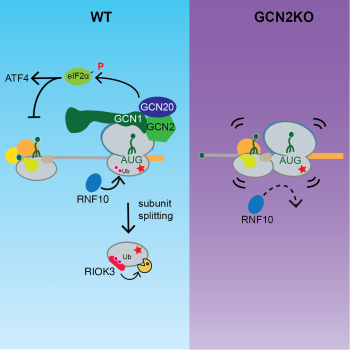
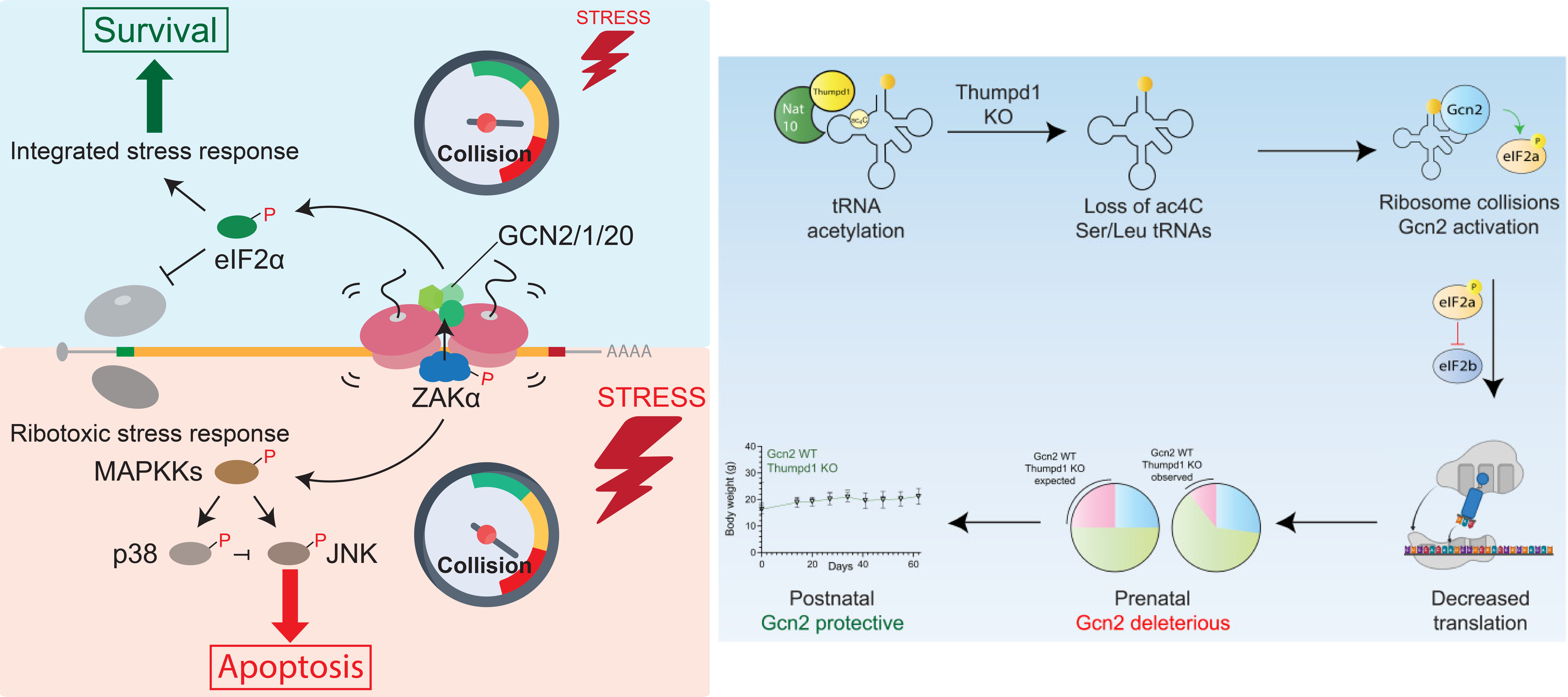
If a stalled ribosome cannot be resolved before a trailing ribosome catches up, ribosome collisions occur. We previously identified that colliding ribosomes serve as a platform that recruits collision-sensing factors and triggers two inter-related but distinct signaling pathways– the ribotoxic stress response and the integrated stress response– to regulate cell fate decisions (Wu et al., Cell 2020). We are actively investigating the molecular mechanisms of this process and its potential implications in normal cell physiology during development (Thalalla Gamage, Khoogar et al., Sci Adv 2025) and in disease-related settings such as cancer.
Additionally, we’re also pursuing how local translation influences cellular stress responses.
Publications
- Bibliography Link
- View Dr. Wu's Complete Bibliography at NCBI.
The integrated stress response regulates 18S nonfunctional rRNA decay in mammals
Transfer RNA acetylation regulates in vivo mammalian stress signaling
Yeast translation elongation factor eEF3 promotes late stages of tRNA translocation.
Ribosome collisions trigger general stress responses to regulate cell fate
Biography

Colin C. C. Wu, Ph.D.
Dr. Wu received his bachelor’s degree from Chiao Tung University in Taiwan, and then his doctoral degree from the Institute of Molecular Biology at Academia Sinica. He undertook his postdoctoral training at Johns Hopkins University, where he became fascinated with RNA biology and the ribosome. During this time, he investigated the molecular mechanisms of translational regulation by the ribosome and became interested in post-transcriptional regulation of gene expression and ribosome homeostasis. He joined the RNA Biology Laboratory at NCI-Frederick as an NIH Stadtman tenure-track investigator in the fall of 2020 to establish the Translational Control of Gene Expression Section.
Job Vacancies
We have no open positions in our group at this time, please check back later.
To see all available positions at CCR, take a look at our Careers page. You can also subscribe to receive CCR's latest job and training opportunities in your inbox.
Team
News
Learn more about CCR research advances, new discoveries and more
on our news section.
Lab Life



Happy Birthday, Jenny!

Lab hike

Fun lunch with the Gonatopoulos-Pournatzis lab